RetrogeneDB ID: | retro_mmus_192 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 18:87756354..87757578(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000069324 | |
| Aliases: | None | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Bhmt | ||
| Ensembl ID: | ENSMUSG00000074768 | ||
| Aliases: | None | ||
| Description: | betaine-homocysteine methyltransferase [Source:MGI Symbol;Acc:MGI:1339972] |

Retrocopy-Parental alignment summary:
>retro_mmus_192
ATGGCACCAGTTGCTGGCAAGAAGGCCAAGAAGGGGATCTTAGAACGCTTAAATGCCGGAGAAGTTGTGATTGGAGATGG
AGGATTTGTCTTTGCACTGGAAAAGAGGGGCTATGTAAAGGCTGGACCCTGGACCCCAGAAGCTGCCGTGGAGCACCCTG
AGGCAGTTCGTCAGCTTCATCGGGAGTTCCTCAGAGCTGGATCGAACGTCATGCAGACCTTCACTTTCTATGCAAGTGAA
GACAAGCTGGAAAACAGAGGAAACTATGTGGCAGAGAAGATTTCTGGGCAGAAAGTCAACGAAGCTGCTTGTGACATTGC
ACGGCAAGTGGCTGATGAAGGAGACGCCTTGGTTGCAGGAGGCGTGAGCCAGACGCCTTCATACCTTAGCTGCAAGAGTG
AGGTAGAAGTGAAAAAGATATTTCGCCAACAGCTAGAGGTGTTCATGAAGAAGAACGTGGACTTCCTCATTGCAGAGTAT
TTTGAACATGTTGAAGAAGCCGTGTGGGCAGTGGAAGCCTTAAAAGCATCTGGTAAGCCCGTAGCAGCTACCATGTGCAT
CGGGCCCGAGGGAGATCTGCATGGCGTGCCCCCTGGAGAGTGTGCCGTGCGTCTGGTGAAAGCAGGCGCCTCCATTGTCG
GCGTGAACTGCCACTTCGACCCCAGCGTCAGCTTACAGACTGTGAAGCTCATGAAGGAGGGTTTGGAGGCTGCGCGGTTG
AAAGCTTACCTGATGAGCCAGCCCCTGGCCTACCATACCCCTGACTGTGGCAAACAGGGATTTATTGATCTCCCAGAATT
CCCCTTTGGATTGGAACCCCGAGTTGCCACTAGATGGGATATTCAAAAATATGCCAGAGAGGCCTACAACCTGGGGGTTA
GGTACATTGGCGGCTGCTGCGGATTTGAGCCCTACCACATCAGGGCGATTGCAGAGGAGTTGGCCCCAGAAAGGGGATTT
TTGCCACCGGCTTCAGAAAAACATGGCAGCTGGGGAAGTGGTTTGGACATGCACACCAAACCCTGGATCAGAGCAAGGGC
CAGAAAGGAATACTGGCAGAATCTGCGAATAGCTTCCGGCAGGCCGTACAACCCTTCCATGTCCCGGCCAGATGCTTGGG
GCGTGACTAAGGGAGCAGCCGAGCTGATGCAGCAGAAGGAGGCCACTACTGAGCAGCAGCTGAGGGAGCTCTTTGAAAAA
CAAAAATTCAAGTCTGCACAGTAG
ATGGCACCAGTTGCTGGCAAGAAGGCCAAGAAGGGGATCTTAGAACGCTTAAATGCCGGAGAAGTTGTGATTGGAGATGG
AGGATTTGTCTTTGCACTGGAAAAGAGGGGCTATGTAAAGGCTGGACCCTGGACCCCAGAAGCTGCCGTGGAGCACCCTG
AGGCAGTTCGTCAGCTTCATCGGGAGTTCCTCAGAGCTGGATCGAACGTCATGCAGACCTTCACTTTCTATGCAAGTGAA
GACAAGCTGGAAAACAGAGGAAACTATGTGGCAGAGAAGATTTCTGGGCAGAAAGTCAACGAAGCTGCTTGTGACATTGC
ACGGCAAGTGGCTGATGAAGGAGACGCCTTGGTTGCAGGAGGCGTGAGCCAGACGCCTTCATACCTTAGCTGCAAGAGTG
AGGTAGAAGTGAAAAAGATATTTCGCCAACAGCTAGAGGTGTTCATGAAGAAGAACGTGGACTTCCTCATTGCAGAGTAT
TTTGAACATGTTGAAGAAGCCGTGTGGGCAGTGGAAGCCTTAAAAGCATCTGGTAAGCCCGTAGCAGCTACCATGTGCAT
CGGGCCCGAGGGAGATCTGCATGGCGTGCCCCCTGGAGAGTGTGCCGTGCGTCTGGTGAAAGCAGGCGCCTCCATTGTCG
GCGTGAACTGCCACTTCGACCCCAGCGTCAGCTTACAGACTGTGAAGCTCATGAAGGAGGGTTTGGAGGCTGCGCGGTTG
AAAGCTTACCTGATGAGCCAGCCCCTGGCCTACCATACCCCTGACTGTGGCAAACAGGGATTTATTGATCTCCCAGAATT
CCCCTTTGGATTGGAACCCCGAGTTGCCACTAGATGGGATATTCAAAAATATGCCAGAGAGGCCTACAACCTGGGGGTTA
GGTACATTGGCGGCTGCTGCGGATTTGAGCCCTACCACATCAGGGCGATTGCAGAGGAGTTGGCCCCAGAAAGGGGATTT
TTGCCACCGGCTTCAGAAAAACATGGCAGCTGGGGAAGTGGTTTGGACATGCACACCAAACCCTGGATCAGAGCAAGGGC
CAGAAAGGAATACTGGCAGAATCTGCGAATAGCTTCCGGCAGGCCGTACAACCCTTCCATGTCCCGGCCAGATGCTTGGG
GCGTGACTAAGGGAGCAGCCGAGCTGATGCAGCAGAAGGAGGCCACTACTGAGCAGCAGCTGAGGGAGCTCTTTGAAAAA
CAAAAATTCAAGTCTGCACAGTAG
ORF - retro_mmus_192 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 100. % |
| Parental protein coverage: | 100. % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MAPVAGKKAKKGILERLNAGEVVIGDGGFVFALEKRGYVKAGPWTPEAAVEHPEAVRQLHREFLRAGSNV |
| MAPVAGKKAKKGILERLNAGEVVIGDGGFVFALEKRGYVKAGPWTPEAAVEHPEAVRQLHREFLRAGSNV | |
| Retrocopy | MAPVAGKKAKKGILERLNAGEVVIGDGGFVFALEKRGYVKAGPWTPEAAVEHPEAVRQLHREFLRAGSNV |
| Parental | MQTFTFYASEDKLENRGNYVAEKISGQKVNEAACDIARQVADEGDALVAGGVSQTPSYLSCKSEVEVKKI |
| MQTFTFYASEDKLENRGNYVAEKISGQKVNEAACDIARQVADEGDALVAGGVSQTPSYLSCKSEVEVKKI | |
| Retrocopy | MQTFTFYASEDKLENRGNYVAEKISGQKVNEAACDIARQVADEGDALVAGGVSQTPSYLSCKSEVEVKKI |
| Parental | FRQQLEVFMKKNVDFLIAEYFEHVEEAVWAVEALKASGKPVAATMCIGPEGDLHGVPPGECAVRLVKAGA |
| FRQQLEVFMKKNVDFLIAEYFEHVEEAVWAVEALKASGKPVAATMCIGPEGDLHGVPPGECAVRLVKAGA | |
| Retrocopy | FRQQLEVFMKKNVDFLIAEYFEHVEEAVWAVEALKASGKPVAATMCIGPEGDLHGVPPGECAVRLVKAGA |
| Parental | SIVGVNCHFDPSVSLQTVKLMKEGLEAARLKAYLMSQPLAYHTPDCGKQGFIDLPEFPFGLEPRVATRWD |
| SIVGVNCHFDPSVSLQTVKLMKEGLEAARLKAYLMSQPLAYHTPDCGKQGFIDLPEFPFGLEPRVATRWD | |
| Retrocopy | SIVGVNCHFDPSVSLQTVKLMKEGLEAARLKAYLMSQPLAYHTPDCGKQGFIDLPEFPFGLEPRVATRWD |
| Parental | IQKYAREAYNLGVRYIGGCCGFEPYHIRAIAEELAPERGFLPPASEKHGSWGSGLDMHTKPWIRARARKE |
| IQKYAREAYNLGVRYIGGCCGFEPYHIRAIAEELAPERGFLPPASEKHGSWGSGLDMHTKPWIRARARKE | |
| Retrocopy | IQKYAREAYNLGVRYIGGCCGFEPYHIRAIAEELAPERGFLPPASEKHGSWGSGLDMHTKPWIRARARKE |
| Parental | YWQNLRIASGRPYNPSMSRPDAWGVTKGAAELMQQKEATTEQQLRELFEKQKFKSAQ |
| YWQNLRIASGRPYNPSMSRPDAWGVTKGAAELMQQKEATTEQQLRELFEKQKFKSAQ | |
| Retrocopy | YWQNLRIASGRPYNPSMSRPDAWGVTKGAAELMQQKEATTEQQLRELFEKQKFKSAQ |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .00 RPM | 0 .09 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 0 .04 RPM |
| SRP007412_heart | 0 .00 RPM | 0 .00 RPM |
| SRP007412_kidney | 1 .12 RPM | 13 .44 RPM |
| SRP007412_liver | 26 .30 RPM | 429 .84 RPM |
| SRP007412_testis | 0 .59 RPM | 7 .68 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_192 was not detected
1 EST(s) were mapped to retro_mmus_192 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| AI530322 | 87756286 | 87756889 | 99.8 | 449 | 1 | 447 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_61924 | 960 libraries | 46 libraries | 17 libraries | 6 libraries | 43 libraries |
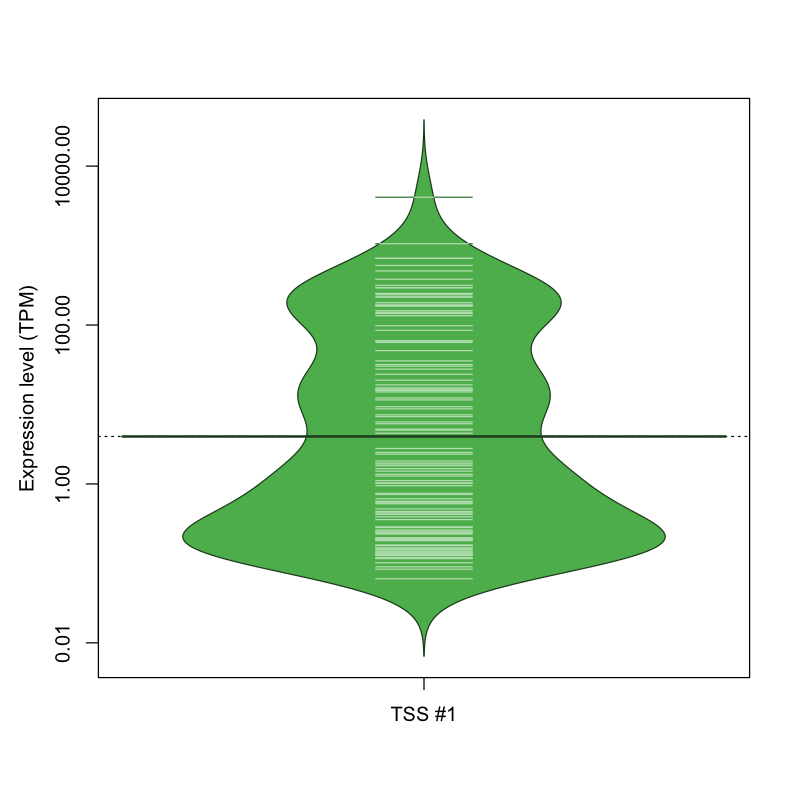
The graphical summary, for retro_mmus_192 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_mmus_192 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_192 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 4 parental genes, and 4 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Ciona intestinalis | ENSCING00000007560 | 1 retrocopy | |
| Monodelphis domestica | ENSMODG00000019726 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000074768 | 1 retrocopy |
retro_mmus_192 ,
|
| Rattus norvegicus | ENSRNOG00000011200 | 1 retrocopy |