RetrogeneDB ID: | retro_mmus_2465 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 4:58483035..58483825(-) | ||
| Located in intron of: | ENSMUSG00000038668 | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000083834 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Rsrc2 | ||
| Ensembl ID: | ENSMUSG00000029422 | ||
| Aliases: | Rsrc2, 1500011J06Rik | ||
| Description: | arginine/serine-rich coiled-coil 2 [Source:MGI Symbol;Acc:MGI:1913489] |

Retrocopy-Parental alignment summary:
>retro_mmus_2465
GTAAGAACCAACTCCTTGTTAAATCAGGCAAGAAGACGTGAATCCAAAGATAAGTCCTCTGGAGACACAAGTCTGAGGAG
CACAATGACAAAGAACATTCTTCCGACAAAGGAAGAGAGCGATTAAATTCATCTGAGAATGGTGAGGACAGACATAAACA
CAAAGAAAGGAAGTCATGATGGGGCAGAAGTCACTCAAGGTCTAGATCGCGTGAAAGGCGCCATCGCAGTAGAAGCAGGG
AAAGGAAGAAGGGATCGGAAAAAAAAATCTCCATCCAGAAGCAGAGACAGGAAGAAGTCAAGATCTAGAAGCAGAGTTAG
AAAACTAAGAATCAGTACACATTCCCAATCAAGATCCATACACAGGCATGGGACTAGAAGCAGGAGCAGATCAAGGAGTC
GAAGTCGAGATAGAAAGAAGAGAATTGAAAAACCAAGAAGATTTAGCAGAAGCCGAAGCCGGACACCTATTCCACCTTCC
TTCAGAGGCAGAAACATTAGCTGGATGCCAAAAAAGCATTAGAAAGGGCAAAGAAATTATAAGAACAACGAGAAAAGGAA
ATGTTGAAAAGCAGAAATAAGAAACTGCTGCAATGGCTGCAGCCACGGGAGGTTCTGTTCTTAATGTCGCTGCAACGTTG
GCATCAGGAATGCAGGTTACTCCTCAGATAACTATGGAAGCTCAAATGGTAGCCCTGCAAGCTAAAGCCTTGGCAGAGAC
TGGCATGGCAGTGCCCACCTACTGTAACCCAGCAGCTGTGAATCTTCGCTGAGCAAAAGAAAAGGAGGAA
GTAAGAACCAACTCCTTGTTAAATCAGGCAAGAAGACGTGAATCCAAAGATAAGTCCTCTGGAGACACAAGTCTGAGGAG
CACAATGACAAAGAACATTCTTCCGACAAAGGAAGAGAGCGATTAAATTCATCTGAGAATGGTGAGGACAGACATAAACA
CAAAGAAAGGAAGTCATGATGGGGCAGAAGTCACTCAAGGTCTAGATCGCGTGAAAGGCGCCATCGCAGTAGAAGCAGGG
AAAGGAAGAAGGGATCGGAAAAAAAAATCTCCATCCAGAAGCAGAGACAGGAAGAAGTCAAGATCTAGAAGCAGAGTTAG
AAAACTAAGAATCAGTACACATTCCCAATCAAGATCCATACACAGGCATGGGACTAGAAGCAGGAGCAGATCAAGGAGTC
GAAGTCGAGATAGAAAGAAGAGAATTGAAAAACCAAGAAGATTTAGCAGAAGCCGAAGCCGGACACCTATTCCACCTTCC
TTCAGAGGCAGAAACATTAGCTGGATGCCAAAAAAGCATTAGAAAGGGCAAAGAAATTATAAGAACAACGAGAAAAGGAA
ATGTTGAAAAGCAGAAATAAGAAACTGCTGCAATGGCTGCAGCCACGGGAGGTTCTGTTCTTAATGTCGCTGCAACGTTG
GCATCAGGAATGCAGGTTACTCCTCAGATAACTATGGAAGCTCAAATGGTAGCCCTGCAAGCTAAAGCCTTGGCAGAGAC
TGGCATGGCAGTGCCCACCTACTGTAACCCAGCAGCTGTGAATCTTCGCTGAGCAAAAGAAAAGGAGGAA
ORF - retro_mmus_2465 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 68.82 % |
| Parental protein coverage: | 72.87 % |
| Number of stop codons detected: | 4 |
| Number of frameshifts detected | 4 |
Retrocopy - Parental Gene Alignment:
| Parental | IRTNFLLKQGRRHESKDKSSK-RHKSEEHNDKEHSSDKGRERLNSSENGEDRHKRKERKSSRGRSHSRSR |
| .RTN.LL.Q.RR.ESKDKSS..RHKSEEHNDKEHSSDKGRERL....N.............R.....RS. | |
| Retrocopy | VRTNSLLNQARRRESKDKSSG<RHKSEEHNDKEHSSDKGRERL----NSSENGEDRHKHKERKS*WGRSH |
| Parental | SRERRHRSRSRERKKSRSRS-RDRKK-SRSRSRDRKKSRSRSRDRKRRIRTRSRSRSRHRHRTRSRSRSR |
| SR.R....R.R.R..SR.R..RDR...S.SRSRDRKKSRSRSR.RK.RI.T.S.SRS.HRH.TRSRSRSR | |
| Retrocopy | SRSRSRERRHRSR--SRERK>RDRXXXSPSRSRDRKKSRSRSRVRKLRISTHSQSRSIHRHGTRSRSRSR |
| Parental | SRSRDRKKRIEKPRRFSRSLSRTPSPPPFRGRNTA-MDAQEALARRLERAKKLQEQREKEM-VEKQKQQE |
| SRSRDRKKRIEKPRRFSRS.SRTP.PP.FRGRN....DA..AL....ERAKKL.EQREKEM.VEKQK..E | |
| Retrocopy | SRSRDRKKRIEKPRRFSRSRSRTPIPPSFRGRNIS<LDAKKAL----ERAKKL*EQREKEM<VEKQK-*E |
| Parental | MAAAAAATGGSVLNVAALLASGTQVTPQIAMAAQMAALQAKALAETGIAVPSYYNPAAVNPMKFAEQEK |
| .AA.AAATGGSVLNVAA.LASG.QVTPQI.M.AQM.ALQAKALAETG.AVP.Y.NPAAVN.....E.E. | |
| Retrocopy | TAAMAAATGGSVLNVAATLASGMQVTPQITMEAQMVALQAKALAETGMAVPTYCNPAAVNLR*AKEKEE |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .02 RPM | 68 .22 RPM |
| SRP007412_cerebellum | 0 .04 RPM | 83 .72 RPM |
| SRP007412_heart | 0 .03 RPM | 39 .54 RPM |
| SRP007412_kidney | 0 .00 RPM | 60 .98 RPM |
| SRP007412_liver | 0 .00 RPM | 33 .13 RPM |
| SRP007412_testis | 0 .14 RPM | 59 .87 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_2465 was not detected
No EST(s) were mapped for retro_mmus_2465 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_105841 | 253 libraries | 357 libraries | 439 libraries | 16 libraries | 7 libraries |
The graphical summary, for retro_mmus_2465 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
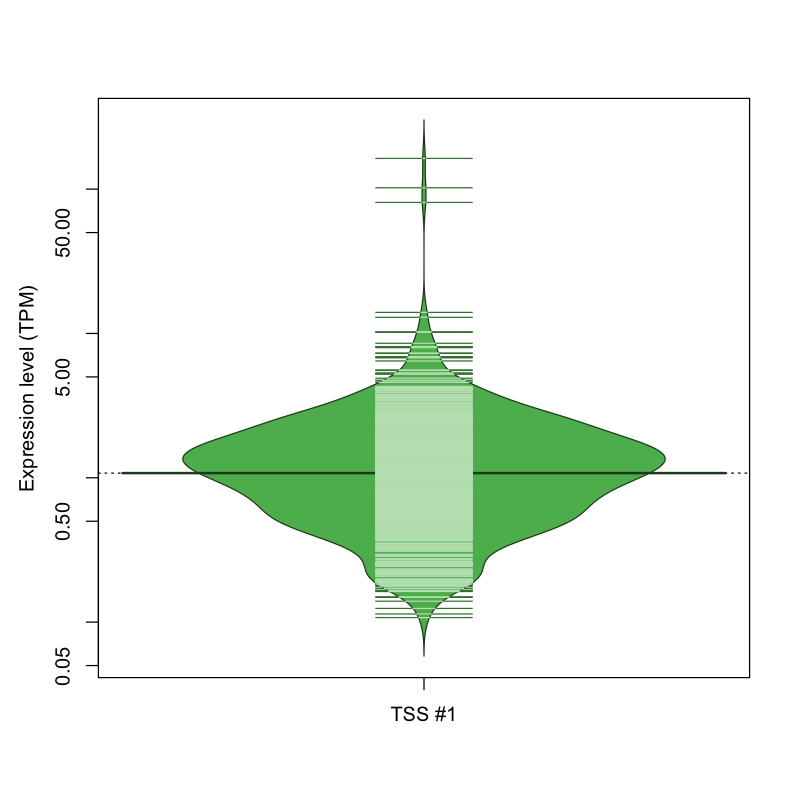
retro_mmus_2465 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_2465 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 5 parental genes, and 5 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Callithrix jacchus | ENSCJAG00000020419 | 1 retrocopy | |
| Dasypus novemcinctus | ENSDNOG00000008912 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000004412 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000029422 | 1 retrocopy |
retro_mmus_2465 ,
|
| Rattus norvegicus | ENSRNOG00000001238 | 1 retrocopy |