RetrogeneDB ID: | retro_hsap_2466 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 20:29873120..29874638(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSG00000224628 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | CDK5RAP3 | ||
| Ensembl ID: | ENSG00000108465 | ||
| Aliases: | CDK5RAP3, C53, HSF-27, IC53, LZAP, MST016, OK/SW-cl.114, PP1553 | ||
| Description: | CDK5 regulatory subunit associated protein 3 [Source:HGNC Symbol;Acc:18673] |

Retrocopy-Parental alignment summary:
>retro_hsap_2466
ATGGAGGACCATCAGCACATGCCCATCGACATCCAGACCAGCAAGCTGCTCGATTGGCTTGTGGACAGAAGGTACTGCAG
CCTGAAATGGCAGAGTCTGGTGCTGACGATCTGAGAGAAGATCAGTGCTGCCATCCAGGACATGCCAGAGAGCGAAGAGA
TCGCCCAGCTGCTGTCTGGATCCTACATTCACTACTTTCACTGCCTAAGAATCCTGGACCTTTTCAAAGGCACAGAGGCC
TCCACGAAAAATATTTTTGGCTGATACTCTTCACAGTGGATGAAGGAATGGCAGGAGATTATAGCCCTGTATGAGAAGGA
CAACACCTACTTAGTGGAACTCTCTAGCCTCCTGGTTTGGAATGTCAACTATGAGATCCCCTCACTGAAGAAGCAGATTG
CCAAGTGCCAGCAGCTGCAGCAAGAATACAGCTGCAAGGAGGAGGAGTGCCAGGCAGGGGCTGCTGAGATGGGGGAGCAG
TTCTACCACTCCTGCAAGCAGTATGGCATCATGGGCGAAAATACCCGAGGAGAACTGCTGGCCCTGGTGAAGGACCTGCC
AAGTCAGCTGGCTGAGACTGGGGCAGCGGCGCAGCAGTCCCTGGGGGAAGCCATCGAAGTGTACCAGGCCTCTGTGGGGT
TTGTGTGTGATAGCCCCACAGAGCAGGTGTTGCCAATGCTGTGGTTCGTGCAGAAGCGGGGAAACTCAACGGTGTACGAG
TGGAGGACAGGGACAGAGCCCTCTGTCGTGGAGCGACCCCACCTCGAGGAGCTTCCTGAGCAGGTGGCAGAAGACGCGAT
TGACTGGGGCGACTTTGGGGTAGAGGCAGTGTCTGGGGGGACTGACGCTGGCATCTCTGCCGAGGCTGCTGGAATCGACT
GGGGCATCTTCCCGGAATCAGACTCAAAGGACCTTGGAGATGATGGGATAGACTAGGGAGACGATGCTGTTGCTTTGCAG
ATCACAGTGGTGGAAGCAGGAATCCAGGCTCCAGAAGGTGTTGCCAGGGGCCCAGATGCTCTGACACTTCTTGAATACAC
TGAGACCCAGAATCAGTTCCTTGATGAGCTCATGGAACTTGAGATCTTCTTAGCCCAGAGAGCAGTGGAGTTGAGTGAGA
AGGCAGATGTCCTGTCTGTGAGCCAGTTCCAGCTGGCTCCAGTCATCCTGCAGGGCCAGACCAAAGAGAAGATGGTTATC
ATGGTGTCAGTGTTGGAAGATCTGATTGGCAAACTTACCAGTCTTCAACTGCAACACCTGTTTATGAACCTGGCCTCACC
AAGGTATGTGGACCGAGTGACTGAATTCCTCCAGCAAAACCTGAAGCAGTCCCAGCTGCTGGCTTTGAAGAAAGAGCTGA
TGGTGCAGAAGCAGCAGGAGGCACTTGAGGAGCAGGAGGCTCTGGAGCCTAAGCTGGACCTGCTGCTGGAGAAGACCAAG
GAGCTGCAGAAGCTGATTGAAGCTGACATCTCCAAGAGGTACAGCGGGCGCCCTGTGAACCTGATGGGAACCTCTCTG
ATGGAGGACCATCAGCACATGCCCATCGACATCCAGACCAGCAAGCTGCTCGATTGGCTTGTGGACAGAAGGTACTGCAG
CCTGAAATGGCAGAGTCTGGTGCTGACGATCTGAGAGAAGATCAGTGCTGCCATCCAGGACATGCCAGAGAGCGAAGAGA
TCGCCCAGCTGCTGTCTGGATCCTACATTCACTACTTTCACTGCCTAAGAATCCTGGACCTTTTCAAAGGCACAGAGGCC
TCCACGAAAAATATTTTTGGCTGATACTCTTCACAGTGGATGAAGGAATGGCAGGAGATTATAGCCCTGTATGAGAAGGA
CAACACCTACTTAGTGGAACTCTCTAGCCTCCTGGTTTGGAATGTCAACTATGAGATCCCCTCACTGAAGAAGCAGATTG
CCAAGTGCCAGCAGCTGCAGCAAGAATACAGCTGCAAGGAGGAGGAGTGCCAGGCAGGGGCTGCTGAGATGGGGGAGCAG
TTCTACCACTCCTGCAAGCAGTATGGCATCATGGGCGAAAATACCCGAGGAGAACTGCTGGCCCTGGTGAAGGACCTGCC
AAGTCAGCTGGCTGAGACTGGGGCAGCGGCGCAGCAGTCCCTGGGGGAAGCCATCGAAGTGTACCAGGCCTCTGTGGGGT
TTGTGTGTGATAGCCCCACAGAGCAGGTGTTGCCAATGCTGTGGTTCGTGCAGAAGCGGGGAAACTCAACGGTGTACGAG
TGGAGGACAGGGACAGAGCCCTCTGTCGTGGAGCGACCCCACCTCGAGGAGCTTCCTGAGCAGGTGGCAGAAGACGCGAT
TGACTGGGGCGACTTTGGGGTAGAGGCAGTGTCTGGGGGGACTGACGCTGGCATCTCTGCCGAGGCTGCTGGAATCGACT
GGGGCATCTTCCCGGAATCAGACTCAAAGGACCTTGGAGATGATGGGATAGACTAGGGAGACGATGCTGTTGCTTTGCAG
ATCACAGTGGTGGAAGCAGGAATCCAGGCTCCAGAAGGTGTTGCCAGGGGCCCAGATGCTCTGACACTTCTTGAATACAC
TGAGACCCAGAATCAGTTCCTTGATGAGCTCATGGAACTTGAGATCTTCTTAGCCCAGAGAGCAGTGGAGTTGAGTGAGA
AGGCAGATGTCCTGTCTGTGAGCCAGTTCCAGCTGGCTCCAGTCATCCTGCAGGGCCAGACCAAAGAGAAGATGGTTATC
ATGGTGTCAGTGTTGGAAGATCTGATTGGCAAACTTACCAGTCTTCAACTGCAACACCTGTTTATGAACCTGGCCTCACC
AAGGTATGTGGACCGAGTGACTGAATTCCTCCAGCAAAACCTGAAGCAGTCCCAGCTGCTGGCTTTGAAGAAAGAGCTGA
TGGTGCAGAAGCAGCAGGAGGCACTTGAGGAGCAGGAGGCTCTGGAGCCTAAGCTGGACCTGCTGCTGGAGAAGACCAAG
GAGCTGCAGAAGCTGATTGAAGCTGACATCTCCAAGAGGTACAGCGGGCGCCCTGTGAACCTGATGGGAACCTCTCTG
ORF - retro_hsap_2466 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 93.87 % |
| Parental protein coverage: | 100. % |
| Number of stop codons detected: | 3 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MEDHQHVPIDIQTSKLLDWLVDRRHCSLKWQSLVLTIREKINAAIQDMPESEEIAQLLSGSYIHYFHCLR |
| MEDHQH.PIDIQTSKLLDWLVDRR.CSLKWQSLVLTI.EKI.AAIQDMPESEEIAQLLSGSYIHYFHCLR | |
| Retrocopy | MEDHQHMPIDIQTSKLLDWLVDRRYCSLKWQSLVLTI*EKISAAIQDMPESEEIAQLLSGSYIHYFHCLR |
| Parental | ILDLLKGTEASTKNIFGRYSSQRMKDWQEIIALYEKDNTYLVELSSLLVRNVNYEIPSLKKQIAKCQQLQ |
| ILDL.KGTEASTKNIFG.YSSQ.MK.WQEIIALYEKDNTYLVELSSLLV.NVNYEIPSLKKQIAKCQQLQ | |
| Retrocopy | ILDLFKGTEASTKNIFG*YSSQWMKEWQEIIALYEKDNTYLVELSSLLVWNVNYEIPSLKKQIAKCQQLQ |
| Parental | QEYSRKEEECQAGAAEMREQFYHSCKQYGITGENVRGELLALVKDLPSQLAEIGAAAQQSLGEAIDVYQA |
| QEYS.KEEECQAGAAEM.EQFYHSCKQYGI.GEN.RGELLALVKDLPSQLAE.GAAAQQSLGEAI.VYQA | |
| Retrocopy | QEYSCKEEECQAGAAEMGEQFYHSCKQYGIMGENTRGELLALVKDLPSQLAETGAAAQQSLGEAIEVYQA |
| Parental | SVGFVCESPTEQVLPMLRFVQKRGNSTVYEWRTGTEPSVVERPHLEELPEQVAEDAIDWGDFGVEAVSEG |
| SVGFVC.SPTEQVLPML.FVQKRGNSTVYEWRTGTEPSVVERPHLEELPEQVAEDAIDWGDFGVEAVS.G | |
| Retrocopy | SVGFVCDSPTEQVLPMLWFVQKRGNSTVYEWRTGTEPSVVERPHLEELPEQVAEDAIDWGDFGVEAVSGG |
| Parental | TDSGISAEAAGIDWGIFPESDSKDPGGDGIDWGDDAVALQITVLEAGTQAPEGVARGPDALTLLEYTETR |
| TD.GISAEAAGIDWGIFPESDSKD.G.DGID.GDDAVALQITV.EAG.QAPEGVARGPDALTLLEYTET. | |
| Retrocopy | TDAGISAEAAGIDWGIFPESDSKDLGDDGID*GDDAVALQITVVEAGIQAPEGVARGPDALTLLEYTETQ |
| Parental | NQFLDELMELEIFLAQRAVELSEEADVLSVSQFQLAPAILQGQTKEKMVTMVSVLEDLIGKLTSLQLQHL |
| NQFLDELMELEIFLAQRAVELSE.ADVLSVSQFQLAP.ILQGQTKEKMV.MVSVLEDLIGKLTSLQLQHL | |
| Retrocopy | NQFLDELMELEIFLAQRAVELSEKADVLSVSQFQLAPVILQGQTKEKMVIMVSVLEDLIGKLTSLQLQHL |
| Parental | FMILASPRYVDRVTEFLQQKLKQSQLLALKKELMVQKQQEALEEQAALEPKLDLLLEKTKELQKLIEADI |
| FM.LASPRYVDRVTEFLQQ.LKQSQLLALKKELMVQKQQEALEEQ.ALEPKLDLLLEKTKELQKLIEADI | |
| Retrocopy | FMNLASPRYVDRVTEFLQQNLKQSQLLALKKELMVQKQQEALEEQEALEPKLDLLLEKTKELQKLIEADI |
| Parental | SKRYSGRPVNLMGTSL |
| SKRYSGRPVNLMGTSL | |
| Retrocopy | SKRYSGRPVNLMGTSL |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .02 RPM | 27 .19 RPM |
| bodymap2_adrenal | 0 .00 RPM | 62 .23 RPM |
| bodymap2_brain | 0 .05 RPM | 26 .89 RPM |
| bodymap2_breast | 0 .04 RPM | 38 .09 RPM |
| bodymap2_colon | 0 .00 RPM | 19 .20 RPM |
| bodymap2_heart | 0 .00 RPM | 10 .76 RPM |
| bodymap2_kidney | 0 .00 RPM | 51 .06 RPM |
| bodymap2_liver | 0 .02 RPM | 31 .22 RPM |
| bodymap2_lung | 0 .00 RPM | 26 .65 RPM |
| bodymap2_lymph_node | 0 .00 RPM | 63 .74 RPM |
| bodymap2_ovary | 0 .00 RPM | 62 .57 RPM |
| bodymap2_prostate | 0 .00 RPM | 45 .78 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 12 .50 RPM |
| bodymap2_testis | 0 .08 RPM | 38 .42 RPM |
| bodymap2_thyroid | 0 .00 RPM | 65 .78 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 24 .89 RPM |
RNA Polymerase II actvity near the 5' end of retro_hsap_2466 was not detected
No EST(s) were mapped for retro_hsap_2466 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_99462 | 196 libraries | 230 libraries | 1192 libraries | 185 libraries | 26 libraries |
The graphical summary, for retro_hsap_2466 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
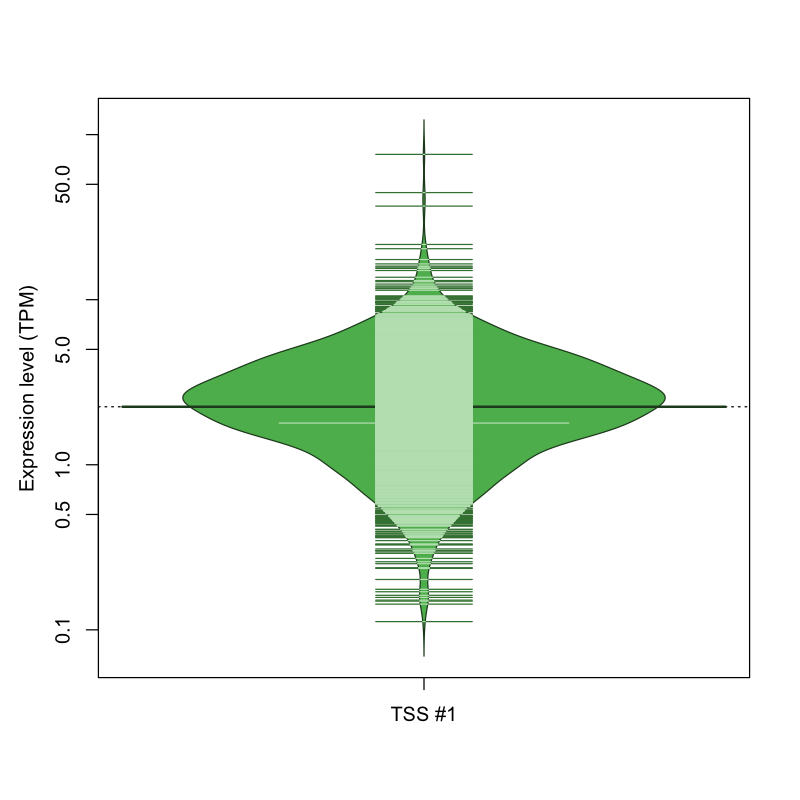
retro_hsap_2466 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_2466 has 2 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Pan troglodytes | retro_ptro_1425 |
| Gorilla gorilla | retro_ggor_1535 |
Parental genes homology:
Parental genes homology involve 6 parental genes, and 7 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Homo sapiens | ENSG00000108465 | 1 retrocopy |
retro_hsap_2466 ,
|
| Gorilla gorilla | ENSGGOG00000022994 | 1 retrocopy | |
| Nomascus leucogenys | ENSNLEG00000002109 | 1 retrocopy | |
| Ochotona princeps | ENSOPRG00000004107 | 2 retrocopies | |
| Pongo abelii | ENSPPYG00000008888 | 1 retrocopy | |
| Pan troglodytes | ENSPTRG00000009345 | 1 retrocopy |
Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .06 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .03 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .00 RPM |
| CEU_NA12760 | 0 .00 RPM |
| CEU_NA12827 | 0 .05 RPM |
| CEU_NA12872 | 0 .08 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .03 RPM |
| FIN_HG00277 | 0 .07 RPM |
| FIN_HG00315 | 0 .03 RPM |
| FIN_HG00321 | 0 .03 RPM |
| FIN_HG00328 | 0 .05 RPM |
| FIN_HG00338 | 0 .04 RPM |
| FIN_HG00349 | 0 .06 RPM |
| FIN_HG00375 | 0 .07 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .02 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .06 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .03 RPM |
| GBR_HG00133 | 0 .05 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .05 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .06 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .03 RPM |
| TSI_NA20532 | 0 .00 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .03 RPM |
| TSI_NA20765 | 0 .00 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .03 RPM |
| TSI_NA20798 | 0 .03 RPM |
| YRI_NA18870 | 0 .07 RPM |
| YRI_NA18907 | 0 .03 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .03 RPM |
| YRI_NA19099 | 0 .03 RPM |
| YRI_NA19114 | 0 .00 RPM |
| YRI_NA19118 | 0 .00 RPM |
| YRI_NA19213 | 0 .05 RPM |
| YRI_NA19214 | 0 .00 RPM |
| YRI_NA19223 | 0 .00 RPM |