RetrogeneDB ID: | retro_hsap_95 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | 14:23511434..23512337(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSG00000222028 | |
| Aliases: | PSMB11, BETA5T | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | PSMB5 | ||
| Ensembl ID: | ENSG00000100804 | ||
| Aliases: | PSMB5, LMPX, MB1, X | ||
| Description: | proteasome (prosome, macropain) subunit, beta type, 5 [Source:HGNC Symbol;Acc:9542] |

Retrocopy-Parental alignment summary:
>retro_hsap_95
ATGGCTCTGCAGGATGTGTGCAAGTGGCAGTCCCCTGACACCCAGGGACCATCACCTCACCTGCCTCGGGCTGGCGGCTG
GGCTGTGCCCCGGGGTTGTGACCCTCAAACCTTCCTGCAGATCCATGGCCCCAGACTGGCCCACGGCACCACCACTCTGG
CCTTCCGCTTCCGTCATGGAGTCATTGCTGCAGCTGACACGCGTTCCTCCTGTGGCAGCTATGTGGCGTGTCCAGCCTCA
TGCAAGGTCATCCCTGTGCACCAGCACCTCCTGGGTACCACCTCTGGCACCTCTGCCGACTGTGCTACCTGGTATCGGGT
ATTACAGCGGGAGCTGCGGCTTCGGGAACTGAGGGAGGGTCAGCTGCCCAGTGTGGCCAGTGCTGCCAAGCTCTTGTCAG
CCATGATGTCTCAATACCGGGGACTGGATCTCTGTGTGGCCACTGCCCTCTGCGGCTGGGACCGCTCTGGCCCTGAGCTC
TTCTACGTCTATAGCGACGGCACCCGCCTGCAGGGGGACATCTTCTCTGTGGGCTCTGGATCTCCCTATGCCTACGGCGT
GCTAGACCGTGGCTATCGCTACGACATGAGCACCCAGGAAGCCTACGCCCTGGCTCGCTGCGCCGTGGCCCACGCCACCC
ACCGTGATGCCTATTCAGGGGGCTCTGTAGACCTTTTCCACGTGCGGGAGAGTGGATGGGAGCATGTGTCACGCAGTGAT
GCCTGTGTGCTGTACGTGGAGTTACAGAAGCTCCTGGAGCCGGAGCCAGAGGAGGATGCCAGCCATGCCCATCCTGAGCC
TGCCACTGCCCACAGAGCTGCAGAAGATAGAGAGCTCTCTGTGGGGCCAGGGGAGGTGACACCAGGAGACTCCAGGATGC
CAGCAGGGACTGAGACGGTGTGA
ATGGCTCTGCAGGATGTGTGCAAGTGGCAGTCCCCTGACACCCAGGGACCATCACCTCACCTGCCTCGGGCTGGCGGCTG
GGCTGTGCCCCGGGGTTGTGACCCTCAAACCTTCCTGCAGATCCATGGCCCCAGACTGGCCCACGGCACCACCACTCTGG
CCTTCCGCTTCCGTCATGGAGTCATTGCTGCAGCTGACACGCGTTCCTCCTGTGGCAGCTATGTGGCGTGTCCAGCCTCA
TGCAAGGTCATCCCTGTGCACCAGCACCTCCTGGGTACCACCTCTGGCACCTCTGCCGACTGTGCTACCTGGTATCGGGT
ATTACAGCGGGAGCTGCGGCTTCGGGAACTGAGGGAGGGTCAGCTGCCCAGTGTGGCCAGTGCTGCCAAGCTCTTGTCAG
CCATGATGTCTCAATACCGGGGACTGGATCTCTGTGTGGCCACTGCCCTCTGCGGCTGGGACCGCTCTGGCCCTGAGCTC
TTCTACGTCTATAGCGACGGCACCCGCCTGCAGGGGGACATCTTCTCTGTGGGCTCTGGATCTCCCTATGCCTACGGCGT
GCTAGACCGTGGCTATCGCTACGACATGAGCACCCAGGAAGCCTACGCCCTGGCTCGCTGCGCCGTGGCCCACGCCACCC
ACCGTGATGCCTATTCAGGGGGCTCTGTAGACCTTTTCCACGTGCGGGAGAGTGGATGGGAGCATGTGTCACGCAGTGAT
GCCTGTGTGCTGTACGTGGAGTTACAGAAGCTCCTGGAGCCGGAGCCAGAGGAGGATGCCAGCCATGCCCATCCTGAGCC
TGCCACTGCCCACAGAGCTGCAGAAGATAGAGAGCTCTCTGTGGGGCCAGGGGAGGTGACACCAGGAGACTCCAGGATGC
CAGCAGGGACTGAGACGGTGTGA
ORF - retro_hsap_95 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 54.68 % |
| Parental protein coverage: | 77.19 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | GIEMLHGTTTLAFKFRHGVIVAADSRATAGAYIASQTVKKVIEINPYLLGTMAGGAADCSFWERLLARQC |
| G....HGTTTLAF.FRHGVI.AAD.R...G.Y.A.....KVI.....LLGT..G..ADC..W.R.L.R.. | |
| Retrocopy | GPRLAHGTTTLAFRFRHGVIAAADTRSSCGSYVACPASCKVIPVHQHLLGTTSGTSADCATWYRVLQREL |
| Parental | RIYELRNKERISVAAASKLLANMVYQYKGMGLSMGTMICGWDKRGPGLYYVDSEGNRISGATFSVGSGSV |
| R..ELR.....SVA.A.KLL..M..QY.G..L...T..CGWD..GP.L.YV.S.G.R..G..FSVGSGS. | |
| Retrocopy | RLRELREGQLPSVASAAKLLSAMMSQYRGLDLCVATALCGWDRSGPELFYVYSDGTRLQGDIFSVGSGSP |
| Parental | YAYGVMDRGYSYDLEVEQAYDLARRAIYQATYRDAYSGGAVNLYHVREDGWIRVSSDNVADLH |
| YAYGV.DRGY.YD.....AY.LAR.A...AT.RDAYSGG.V.L.HVRE.GW..VS......L. | |
| Retrocopy | YAYGVLDRGYRYDMSTQEAYALARCAVAHATHRDAYSGGSVDLFHVRESGWEHVSRSDACVLY |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .02 RPM | 40 .33 RPM |
| bodymap2_adrenal | 0 .00 RPM | 51 .87 RPM |
| bodymap2_brain | 0 .00 RPM | 57 .17 RPM |
| bodymap2_breast | 0 .00 RPM | 54 .05 RPM |
| bodymap2_colon | 0 .00 RPM | 53 .00 RPM |
| bodymap2_heart | 0 .00 RPM | 43 .48 RPM |
| bodymap2_kidney | 0 .00 RPM | 62 .48 RPM |
| bodymap2_liver | 0 .00 RPM | 56 .84 RPM |
| bodymap2_lung | 0 .05 RPM | 38 .98 RPM |
| bodymap2_lymph_node | 0 .05 RPM | 43 .25 RPM |
| bodymap2_ovary | 0 .00 RPM | 45 .09 RPM |
| bodymap2_prostate | 0 .00 RPM | 52 .24 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 74 .26 RPM |
| bodymap2_testis | 0 .02 RPM | 48 .66 RPM |
| bodymap2_thyroid | 0 .00 RPM | 67 .10 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 18 .13 RPM |
RNA Polymerase II actvity near the 5' end of retro_hsap_95 was not detected
10 EST(s) were mapped to retro_hsap_95 retrocopy
| EST ID | Start | End | Identity | Match | Mis-match | Score |
|---|---|---|---|---|---|---|
| EG357703 | 23511446 | 23511822 | 97.9 | 368 | 8 | 360 |
| EG357704 | 23511361 | 23511768 | 99.3 | 407 | 0 | 406 |
| HY065825 | 23511446 | 23511888 | 99.8 | 441 | 1 | 440 |
| HY081305 | 23511420 | 23511917 | 99.8 | 495 | 1 | 494 |
| HY082318 | 23511420 | 23511893 | 99.8 | 472 | 1 | 471 |
| HY082435 | 23511420 | 23511868 | 99.4 | 444 | 3 | 441 |
| HY084301 | 23511444 | 23511899 | 100 | 455 | 0 | 455 |
| HY097430 | 23511417 | 23511892 | 100 | 475 | 0 | 475 |
| HY102966 | 23511478 | 23511922 | 99.1 | 440 | 4 | 436 |
| HY102983 | 23511420 | 23511863 | 98.2 | 440 | 2 | 435 |
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_36978 | 1820 libraries | 5 libraries | 1 library | 1 library | 2 libraries |
The graphical summary, for retro_hsap_95 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
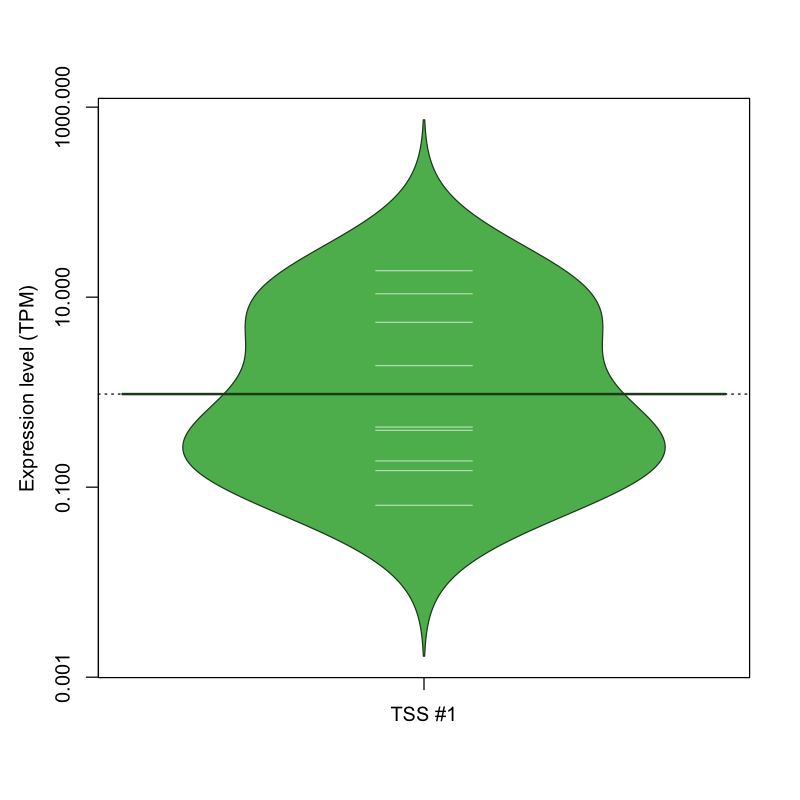
retro_hsap_95 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_95 has 9 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 11 parental genes, and 16 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Callithrix jacchus | ENSCJAG00000008299 | 4 retrocopies | |
| Homo sapiens | ENSG00000100804 | 2 retrocopies |
retro_hsap_1634, retro_hsap_95 ,
|
| Latimeria chalumnae | ENSLACG00000015194 | 1 retrocopy | |
| Macropus eugenii | ENSMEUG00000003572 | 1 retrocopy | |
| Myotis lucifugus | ENSMLUG00000014281 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000006007 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000022193 | 1 retrocopy | |
| Nomascus leucogenys | ENSNLEG00000014891 | 2 retrocopies | |
| Oryctolagus cuniculus | ENSOCUG00000013109 | 1 retrocopy | |
| Pongo abelii | ENSPPYG00000005654 | 1 retrocopy | |
| Sus scrofa | ENSSSCG00000002036 | 1 retrocopy |
Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .00 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .00 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .00 RPM |
| CEU_NA12760 | 0 .00 RPM |
| CEU_NA12827 | 0 .05 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .00 RPM |
| FIN_HG00277 | 0 .00 RPM |
| FIN_HG00315 | 0 .00 RPM |
| FIN_HG00321 | 0 .00 RPM |
| FIN_HG00328 | 0 .00 RPM |
| FIN_HG00338 | 0 .00 RPM |
| FIN_HG00349 | 0 .00 RPM |
| FIN_HG00375 | 0 .00 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .00 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .00 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .00 RPM |
| GBR_HG00133 | 0 .02 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .00 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .00 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .00 RPM |
| TSI_NA20532 | 0 .00 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .00 RPM |
| TSI_NA20765 | 0 .00 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .00 RPM |
| TSI_NA20798 | 0 .00 RPM |
| YRI_NA18870 | 0 .00 RPM |
| YRI_NA18907 | 0 .00 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .00 RPM |
| YRI_NA19099 | 0 .00 RPM |
| YRI_NA19114 | 0 .00 RPM |
| YRI_NA19118 | 0 .00 RPM |
| YRI_NA19213 | 0 .02 RPM |
| YRI_NA19214 | 0 .02 RPM |
| YRI_NA19223 | 0 .00 RPM |