RetrogeneDB ID: | retro_mmus_1072 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 13:3138787..3140112(-) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | None | |
| Aliases: | None | ||
| Status: | NOVEL | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Ptbp1 | ||
| Ensembl ID: | ENSMUSG00000006498 | ||
| Aliases: | Ptbp1, AA407203, AL033359, HNRPI, PTB-1, PTB2, PTB3, PTB4, Ptb, pPTB | ||
| Description: | polypyrimidine tract binding protein 1 [Source:MGI Symbol;Acc:MGI:97791] |

Retrocopy-Parental alignment summary:
>retro_mmus_1072
ATGGACGGCATCGTCCCAGACATAGCAGTCGGTACAAAGCAGGGATCCGACGAGCTCTTCTCCACGTGTGTCAGCAACGG
CCCCTTCATCATGAGCAGCTCTGCCTCAGCAGCCAATGGAAACGATAGCAAGAAGTTCAAAGGTGACAACAGGAGCGCAG
GAGTCCCTTCCAGAGTCATCCACGTCAGAAAGCTGCCCAGCGATGTCACTGAGGGCGAGGTCATCTCCCTAGGGCTGCCC
TTTGGAAAGGTTACCAACCTTCTTATGCTGAAGGGGAAGAACCAGGCCTTCACTGAAATGAACACAGAGGAGACTGCCAA
CACTATGGTTAACTACTATACATCGGTAGCGCCAGTGCTTCGTGGACAGCCCATCTACATCCAGTTCTCCAACCACAAAG
AGCTCGAGACCGACAGCTCGCCCAACCAGGCACGTGCCCAGGCAGCCCTGCAGGCTGTAAACTCTGTCCATTCTGGAAAC
CTGGCCTTGGCAGCGTCTGCTGCTGCCGTAGACGCAGGAATGGCAATGGCAGGGCAGAGTCCAGCGCTCAGGATCATTGT
CGAAAACCTTTTCTACCCAGTGACCCTGGACATGGTGCACCAGATCTTCTCTAAGTTTGGCACCGTCCTGAAGATCATCA
CATTCACCAAGAACAACCAGTTCCAGGCGCTGCTGCAGTATGCTGACCCTGTGAGTGCCCAGCATGCCAAGCTGTCCCTG
GATGGCTAGAACATCTACAACGCCTGCTGCACGCTACGCATTGACTTCTCCAAGCTCACCAGTCTCAACATCAAATACAA
CAATGATAAGAGCAGAGACTACACTCGACCTGACCTGCCATCTGGAGACAGCCAGCCTTCACTAGACCAGACCATGGCAG
CAGCCTTTGGTGCGCCCGGCATAATGTCAGCCTCTCCATATGCAGGAGCCCGGTTCATCCCACCTCAGGCCGCAGGCCTC
TCTGTCCCTAATGTCCATGGAGCCTTGGCTCCCCTGGCCATCTCGTCTGCTGCTGCTGCTGCTGCGGCCAGCTGCATTGC
CATCCCAGGGTTGGCAGGTGCTGGGAATTCCGTCCTTTTGGTCAGCAACCTGAACCCTGAGATAGTCACACCCCAAAGCC
TCTTTATTCTCTTCGGCGTCTACGGTGATGTGCAGCGGGTGAAGATCCTGTTCAATAAGGAGAAGCGCTTGTGCAGATGG
CAGACGGCAGCCAGGCCCAGCTGGCCATGAGCCACATGAACGGGCACAAGCACGGGAAGTCAGTGCGCATCACACTGTCC
AAGCATCAGAGTGTGCAGCTGCCTCGGGAGGGTCAGGAGGACCAG
ATGGACGGCATCGTCCCAGACATAGCAGTCGGTACAAAGCAGGGATCCGACGAGCTCTTCTCCACGTGTGTCAGCAACGG
CCCCTTCATCATGAGCAGCTCTGCCTCAGCAGCCAATGGAAACGATAGCAAGAAGTTCAAAGGTGACAACAGGAGCGCAG
GAGTCCCTTCCAGAGTCATCCACGTCAGAAAGCTGCCCAGCGATGTCACTGAGGGCGAGGTCATCTCCCTAGGGCTGCCC
TTTGGAAAGGTTACCAACCTTCTTATGCTGAAGGGGAAGAACCAGGCCTTCACTGAAATGAACACAGAGGAGACTGCCAA
CACTATGGTTAACTACTATACATCGGTAGCGCCAGTGCTTCGTGGACAGCCCATCTACATCCAGTTCTCCAACCACAAAG
AGCTCGAGACCGACAGCTCGCCCAACCAGGCACGTGCCCAGGCAGCCCTGCAGGCTGTAAACTCTGTCCATTCTGGAAAC
CTGGCCTTGGCAGCGTCTGCTGCTGCCGTAGACGCAGGAATGGCAATGGCAGGGCAGAGTCCAGCGCTCAGGATCATTGT
CGAAAACCTTTTCTACCCAGTGACCCTGGACATGGTGCACCAGATCTTCTCTAAGTTTGGCACCGTCCTGAAGATCATCA
CATTCACCAAGAACAACCAGTTCCAGGCGCTGCTGCAGTATGCTGACCCTGTGAGTGCCCAGCATGCCAAGCTGTCCCTG
GATGGCTAGAACATCTACAACGCCTGCTGCACGCTACGCATTGACTTCTCCAAGCTCACCAGTCTCAACATCAAATACAA
CAATGATAAGAGCAGAGACTACACTCGACCTGACCTGCCATCTGGAGACAGCCAGCCTTCACTAGACCAGACCATGGCAG
CAGCCTTTGGTGCGCCCGGCATAATGTCAGCCTCTCCATATGCAGGAGCCCGGTTCATCCCACCTCAGGCCGCAGGCCTC
TCTGTCCCTAATGTCCATGGAGCCTTGGCTCCCCTGGCCATCTCGTCTGCTGCTGCTGCTGCTGCGGCCAGCTGCATTGC
CATCCCAGGGTTGGCAGGTGCTGGGAATTCCGTCCTTTTGGTCAGCAACCTGAACCCTGAGATAGTCACACCCCAAAGCC
TCTTTATTCTCTTCGGCGTCTACGGTGATGTGCAGCGGGTGAAGATCCTGTTCAATAAGGAGAAGCGCTTGTGCAGATGG
CAGACGGCAGCCAGGCCCAGCTGGCCATGAGCCACATGAACGGGCACAAGCACGGGAAGTCAGTGCGCATCACACTGTCC
AAGCATCAGAGTGTGCAGCTGCCTCGGGAGGGTCAGGAGGACCAG
ORF - retro_mmus_1072 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 93.1 % |
| Parental protein coverage: | 80.72 % |
| Number of stop codons detected: | 1 |
| Number of frameshifts detected | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | MDGIVPDIAVGTKRGSDELFSTCVSNGPFIMSSSASAANGNDSKKFKGDNRSAGVPSRVIHVRKLPSDVT |
| MDGIVPDIAVGTK.GSDELFSTCVSNGPFIMSSSASAANGNDSKKFKGDNRSAGVPSRVIHVRKLPSDVT | |
| Retrocopy | MDGIVPDIAVGTKQGSDELFSTCVSNGPFIMSSSASAANGNDSKKFKGDNRSAGVPSRVIHVRKLPSDVT |
| Parental | EGEVISLGLPFGKVTNLLMLKGKNQAFIEMNTEEAANTMVNYYTSVAPVLRGQPIYIQFSNHKELKTDSS |
| EGEVISLGLPFGKVTNLLMLKGKNQAF.EMNTEE.ANTMVNYYTSVAPVLRGQPIYIQFSNHKEL.TDSS | |
| Retrocopy | EGEVISLGLPFGKVTNLLMLKGKNQAFTEMNTEETANTMVNYYTSVAPVLRGQPIYIQFSNHKELETDSS |
| Parental | PNQARAQAALQAVNSVQSGNLALAASAAAVDAGMAMAGQSPVLRIIVENLFYPVTLDVLHQIFSKFGTVL |
| PNQARAQAALQAVNSV.SGNLALAASAAAVDAGMAMAGQSP.LRIIVENLFYPVTLD..HQIFSKFGTVL | |
| Retrocopy | PNQARAQAALQAVNSVHSGNLALAASAAAVDAGMAMAGQSPALRIIVENLFYPVTLDMVHQIFSKFGTVL |
| Parental | KIITFTKNNQFQALLQYADPVSAQHAKLSLDGQNIYNACCTLRIDFSKLTSLNVKYNNDKSRDYTRPDLP |
| KIITFTKNNQFQALLQYADPVSAQHAKLSLDG.NIYNACCTLRIDFSKLTSLN.KYNNDKSRDYTRPDLP | |
| Retrocopy | KIITFTKNNQFQALLQYADPVSAQHAKLSLDG*NIYNACCTLRIDFSKLTSLNIKYNNDKSRDYTRPDLP |
| Parental | SGDSQPSLDQTMAAAFGAPGIMSASPYAGAGFPPTFAIPQAAGLSVPNVHGALAPLAIPSAAAAAAASRI |
| SGDSQPSLDQTMAAAFGAPGIMSASPYAGA.F.P....PQAAGLSVPNVHGALAPLAI........AS.I | |
| Retrocopy | SGDSQPSLDQTMAAAFGAPGIMSASPYAGARFIP----PQAAGLSVPNVHGALAPLAISXXXXXXXASCI |
| Parental | AIPGLAGAGNSVLLVSNLNPERVTPQSLFILFGVYGDVQRVKILFNKKEN-ALVQMADGSQAQLAMSHLN |
| AIPGLAGAGNSVLLVSNLNPE.VTPQSLFILFGVYGDVQRVKILFN.KE..ALVQMADGSQAQLAMSH.N | |
| Retrocopy | AIPGLAGAGNSVLLVSNLNPEIVTPQSLFILFGVYGDVQRVKILFN-KEK<ALVQMADGSQAQLAMSHMN |
| Parental | GHKLHGKSVRITLSKHQSVQLPREGQEDQ |
| GHK.HGKSVRITLSKHQSVQLPREGQEDQ | |
| Retrocopy | GHK-HGKSVRITLSKHQSVQLPREGQEDQ |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .05 RPM | 22 .78 RPM |
| SRP007412_cerebellum | 0 .00 RPM | 31 .78 RPM |
| SRP007412_heart | 0 .06 RPM | 33 .18 RPM |
| SRP007412_kidney | 0 .04 RPM | 81 .75 RPM |
| SRP007412_liver | 0 .11 RPM | 63 .82 RPM |
| SRP007412_testis | 5 .72 RPM | 68 .52 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_1072 was not detected
No EST(s) were mapped for retro_mmus_1072 retrocopy.
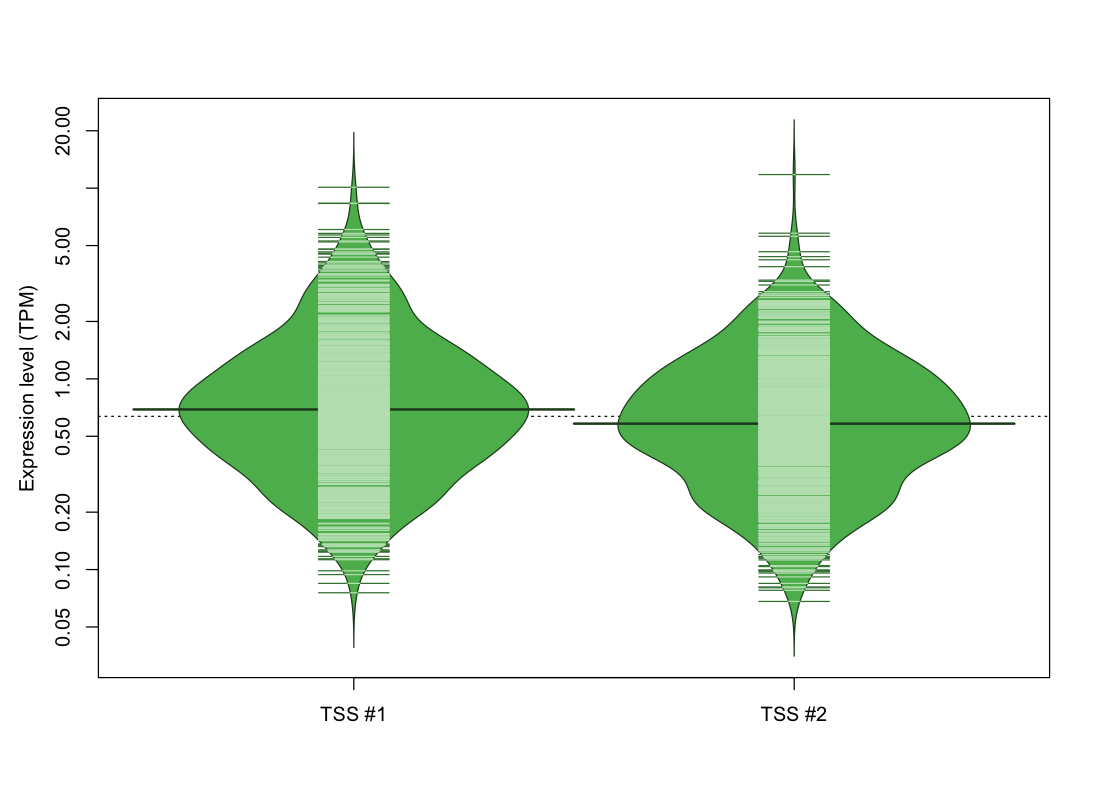
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_28389 | 285 libraries | 538 libraries | 240 libraries | 8 libraries | 1 library |
| TSS #2 | TSS_28390 | 330 libraries | 551 libraries | 188 libraries | 2 libraries | 1 library |
The graphical summary, for retro_mmus_1072 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
retro_mmus_1072 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_1072 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 7 parental genes, and 9 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Ciona savignyi | ENSCSAVG00000007421 | 1 retrocopy | |
| Equus caballus | ENSECAG00000018845 | 1 retrocopy | |
| Echinops telfairi | ENSETEG00000006890 | 1 retrocopy | |
| Macropus eugenii | ENSMEUG00000006710 | 2 retrocopies | |
| Myotis lucifugus | ENSMLUG00000003320 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000006498 | 1 retrocopy |
retro_mmus_1072 ,
|
| Rattus norvegicus | ENSRNOG00000010448 | 2 retrocopies |