RetrogeneDB ID: | retro_hsap_50 | ||
Retrocopylocation | Organism: | Human (Homo sapiens) | |
| Coordinates: | X:73524101..73524602(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSG00000187969 | |
| Aliases: | ZCCHC13, CNBP2, ZNF9L | ||
| Status: | KNOWN_PROTEIN_CODING | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | CNBP | ||
| Ensembl ID: | ENSG00000169714 | ||
| Aliases: | CNBP, CNBP1, DM2, PROMM, RNF163, ZCCHC22, ZNF9 | ||
| Description: | CCHC-type zinc finger, nucleic acid binding protein [Source:HGNC Symbol;Acc:13164] |

Retrocopy-Parental alignment summary:
>retro_hsap_50
ATGAGCAGTAAGGATTTCTTCGCGTGTGGACACTCTGGCCATTGGGCTCGGGGATGCCCTAGAGGAGGAGCTGGAGGGCG
AAGAGGTGGAGGCCATGGCAGAGGTTCTCAATGTGGTTCCACCACCCTATCTTACACCTGTTACTGCTGTGGTGAGTCCG
GTCGTAATGCTAAGAACTGTGTCCTTCTCGGAAACATCTGCTACAACTGTGGGAGAAGCGGCCACATCGCCAAAGACTGT
AAGGATCCTAAACGAGAGAGACGCCAACACTGTTATACCTGCGGCAGACTAGGACATCTGGCTCGTGACTGTGATCGTCA
GAAAGAGCAGAAATGCTACTCTTGCGGCAAACTTGGGCACATTCAGAAAGACTGCGCCCAGGTCAAGTGTTACCGATGCG
GCGAGATTGGCCACGTGGCCATCAATTGCAGCAAGGCGAGGCCAGGTCAACTGCTACCGCTGCGGCAAATCCCGACATCT
AGCCAAGGAATGTCCCAGTGA
ATGAGCAGTAAGGATTTCTTCGCGTGTGGACACTCTGGCCATTGGGCTCGGGGATGCCCTAGAGGAGGAGCTGGAGGGCG
AAGAGGTGGAGGCCATGGCAGAGGTTCTCAATGTGGTTCCACCACCCTATCTTACACCTGTTACTGCTGTGGTGAGTCCG
GTCGTAATGCTAAGAACTGTGTCCTTCTCGGAAACATCTGCTACAACTGTGGGAGAAGCGGCCACATCGCCAAAGACTGT
AAGGATCCTAAACGAGAGAGACGCCAACACTGTTATACCTGCGGCAGACTAGGACATCTGGCTCGTGACTGTGATCGTCA
GAAAGAGCAGAAATGCTACTCTTGCGGCAAACTTGGGCACATTCAGAAAGACTGCGCCCAGGTCAAGTGTTACCGATGCG
GCGAGATTGGCCACGTGGCCATCAATTGCAGCAAGGCGAGGCCAGGTCAACTGCTACCGCTGCGGCAAATCCCGACATCT
AGCCAAGGAATGTCCCAGTGA
ORF - retro_hsap_50 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 68.28 % |
| Parental protein coverage: | 85.29 % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MSSNECFKCGRSGHWARECPTGGGRGRGMRSRGRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDACYNC |
| MSS...F.CG.SGHWAR.CP.GG..GR.....GRG.Q..S..L...CY.CGESG..AK.C.L....CYNC | |
| Retrocopy | MSSKDFFACGHSGHWARGCPRGGAGGRRGGGHGRGSQCGSTTLSYTCYCCGESGRNAKNCVLLGNICYNC |
| Parental | GRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHVA |
| GR.GHIAKDCK.PKRER.Q.CY.CG..GHLARDCD...EQKCYSCG..GHIQKDC..VKCYRCGE.GHVA | |
| Retrocopy | GRSGHIAKDCKDPKRERRQHCYTCGRLGHLARDCDRQKEQKCYSCGKLGHIQKDCAQVKCYRCGEIGHVA |
| Parental | INCSK |
| INCSK | |
| Retrocopy | INCSK |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .00 RPM | 341 .52 RPM |
| bodymap2_adrenal | 0 .00 RPM | 419 .42 RPM |
| bodymap2_brain | 0 .00 RPM | 307 .18 RPM |
| bodymap2_breast | 0 .00 RPM | 305 .86 RPM |
| bodymap2_colon | 0 .00 RPM | 486 .35 RPM |
| bodymap2_heart | 0 .00 RPM | 390 .77 RPM |
| bodymap2_kidney | 0 .00 RPM | 225 .13 RPM |
| bodymap2_liver | 0 .00 RPM | 210 .60 RPM |
| bodymap2_lung | 0 .00 RPM | 606 .40 RPM |
| bodymap2_lymph_node | 0 .00 RPM | 353 .87 RPM |
| bodymap2_ovary | 0 .00 RPM | 413 .37 RPM |
| bodymap2_prostate | 0 .00 RPM | 608 .89 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 778 .15 RPM |
| bodymap2_testis | 4 .94 RPM | 379 .46 RPM |
| bodymap2_thyroid | 0 .00 RPM | 497 .30 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 495 .78 RPM |
RNA Polymerase II actvity near the 5' end of retro_hsap_50 was not detected
No EST(s) were mapped for retro_hsap_50 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_201196 | 1825 libraries | 1 library | 1 library | 2 libraries | 0 libraries |
| TSS #2 | TSS_201197 | 1824 libraries | 2 libraries | 1 library | 2 libraries | 0 libraries |
The graphical summary, for retro_hsap_50 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
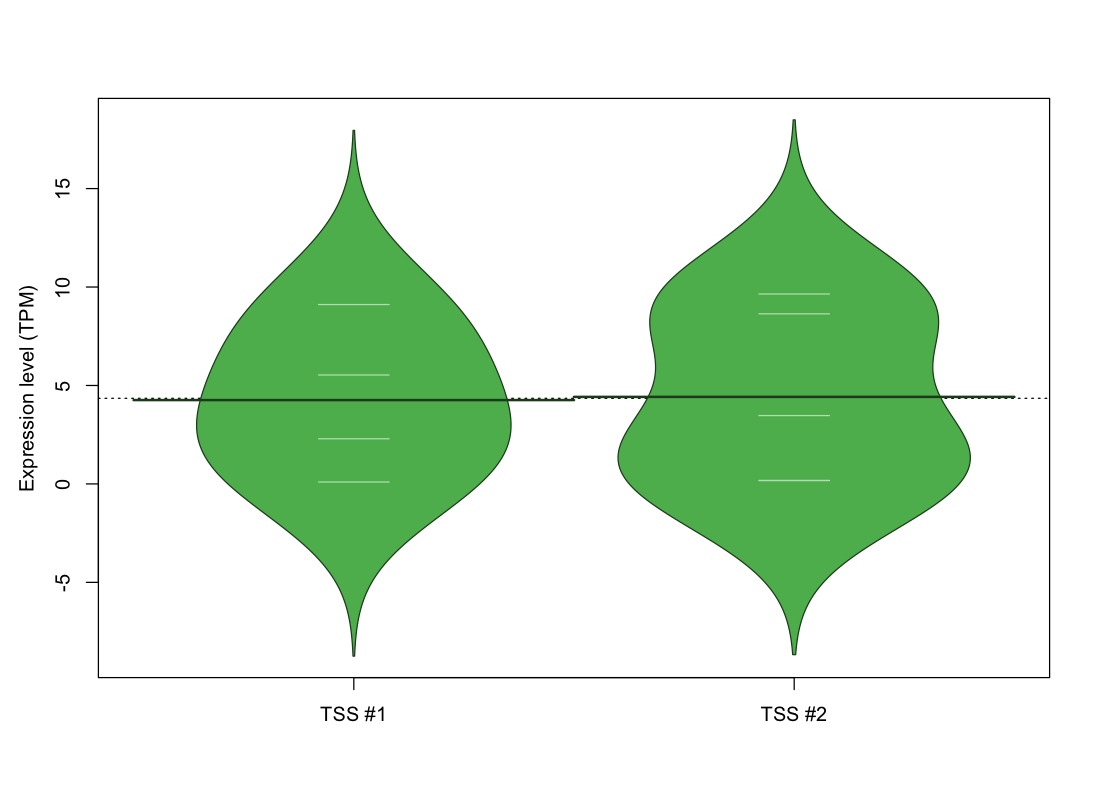
retro_hsap_50 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_50 has 2 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Pongo abelii | retro_pabe_3658 |
| Equus caballus | retro_ecab_1072 |
Parental genes homology:
Parental genes homology involve 22 parental genes, and 37 retrocopies.Expression level across human populations :
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .00 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .00 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .00 RPM |
| CEU_NA12760 | 0 .00 RPM |
| CEU_NA12827 | 0 .00 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .00 RPM |
| FIN_HG00277 | 0 .00 RPM |
| FIN_HG00315 | 0 .00 RPM |
| FIN_HG00321 | 0 .00 RPM |
| FIN_HG00328 | 0 .00 RPM |
| FIN_HG00338 | 0 .00 RPM |
| FIN_HG00349 | 0 .00 RPM |
| FIN_HG00375 | 0 .00 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .00 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .00 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .00 RPM |
| GBR_HG00133 | 0 .00 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .00 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .00 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .00 RPM |
| TSI_NA20532 | 0 .00 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .00 RPM |
| TSI_NA20765 | 0 .00 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .00 RPM |
| TSI_NA20798 | 0 .00 RPM |
| YRI_NA18870 | 0 .00 RPM |
| YRI_NA18907 | 0 .00 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .00 RPM |
| YRI_NA19099 | 0 .00 RPM |
| YRI_NA19114 | 0 .00 RPM |
| YRI_NA19118 | 0 .00 RPM |
| YRI_NA19213 | 0 .00 RPM |
| YRI_NA19214 | 0 .00 RPM |
| YRI_NA19223 | 0 .00 RPM |