RetrogeneDB ID: | retro_mmus_2588 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 5:95974239..95975623(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | None | |
| Aliases: | None | ||
| Status: | NOVEL | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Odc1 | ||
| Ensembl ID: | ENSMUSG00000011179 | ||
| Aliases: | Odc1, ODC | ||
| Description: | ornithine decarboxylase, structural 1 [Source:MGI Symbol;Acc:MGI:97402] |

Retrocopy-Parental alignment summary:
>retro_mmus_2588
ATGAGCAACTTTACTAAGGACGAGTTTGACTGCCACATCCTTGATGAAGGCTTTACTGCTAAGGACATTCTGGAACAAAA
AAATCAATGAAGTCTCTTCCTCTGACGATAAGGATGCGTTCTATGTTGCGGACCTCGGAGACATTCTAAAGAAGCATCTG
AGGTGGCTAAAAGCTCTTCCCCGCGTTACTCCCGTTTACGCAGTCAAGTGTAACGATAGCAGAGCCATAGTGAGCACCCT
AGCTGCCTTTGGGATAGGATTTGACTGTGCAAGCAAGACTGAAATACAGTTGGTACAGGGGCCTGGGGTGCCTCCAGAGA
CGATTATCTATGCAAATCCTTGTAAACAAGTCTCTCAAATCAAGTATGCTGCCAGTAGCGGAGTCCAGATGATGACTTTT
GACAGTGAAATTGAACTGATGAAAGTCGCCAGAGCACATCCAAAGGCAAAGTTGGTTCTGCGGATTGCCACTGATGATTC
CAAAGCTGTCTGTCGCCTCAGTGTTAAGTTTGGTGCCACACTCAAAACCAGCAGGCTTCTCTTGGAACGGGCAAAAGAGC
TAAATATTGACGTCATTGGTGTGAGCTTCCATGTGGGCAGTGGATGTACTGATCCTGAGACCTTCGTGCAGGCAGTGTCG
GATGCCCGCTGTGTGTTTGACATGGCAACAGAAGTTGGTTTCAGCATGCATCTGCTTGATATTGGTGGTGGCTTTCCTGG
ATCTGAAGATACAAAGCTTAAATTTGAAGAGATCACCAGTGTAATCAAACCAGCTCTGGACAAGTACTTCCCATCAGACT
CTGGAGTGAGAATCATAGCCGAGCCAGGCAGCTTTCACGCTTGCAGTCAACATCATTGCCAAAAAAAAAAAAAAAAAAAA
AAAAACGTGTGAAAGGAGCAGCCGAGTTCTGACGATGATGATGAGTCAAATGAGCAAACCTTCATGTATTATGTGAATGA
TGGAGTGTACGGATCATTTAACTGCATTCTTTATGACCATGCCCATGTGAAGGCCCTGCTGCAGAAGAGACCCAAGCCAG
ATGAGAAGTATTACTCATCCAGCATCTGGGGACCAACATGTGATGGCCTTGATCGGATCCTGGAGCGCTGTAACCTGCCT
GAAATGCATGTGGGTGATTGGATGCTGTTTGAGAACATGGGTGCATACACCGTTGCTGCTGCTTCTACTTTCAATGGGTT
CCAGAGGCCAAACATCTACTATGTAATGTCACGGCCAATGTGGCAACTCATGAAACAGATCCAGAGCCATGGCTTCCCGC
CGGAGGTGGAGGAGCAGGATGATGGCACGCTGCCCATGTCTTGTGCCCAGGAGAGCGGGATGGACCGTCACCCTGCAGCC
TGTGCTTCTGCTAGGATCAATGTG
ATGAGCAACTTTACTAAGGACGAGTTTGACTGCCACATCCTTGATGAAGGCTTTACTGCTAAGGACATTCTGGAACAAAA
AAATCAATGAAGTCTCTTCCTCTGACGATAAGGATGCGTTCTATGTTGCGGACCTCGGAGACATTCTAAAGAAGCATCTG
AGGTGGCTAAAAGCTCTTCCCCGCGTTACTCCCGTTTACGCAGTCAAGTGTAACGATAGCAGAGCCATAGTGAGCACCCT
AGCTGCCTTTGGGATAGGATTTGACTGTGCAAGCAAGACTGAAATACAGTTGGTACAGGGGCCTGGGGTGCCTCCAGAGA
CGATTATCTATGCAAATCCTTGTAAACAAGTCTCTCAAATCAAGTATGCTGCCAGTAGCGGAGTCCAGATGATGACTTTT
GACAGTGAAATTGAACTGATGAAAGTCGCCAGAGCACATCCAAAGGCAAAGTTGGTTCTGCGGATTGCCACTGATGATTC
CAAAGCTGTCTGTCGCCTCAGTGTTAAGTTTGGTGCCACACTCAAAACCAGCAGGCTTCTCTTGGAACGGGCAAAAGAGC
TAAATATTGACGTCATTGGTGTGAGCTTCCATGTGGGCAGTGGATGTACTGATCCTGAGACCTTCGTGCAGGCAGTGTCG
GATGCCCGCTGTGTGTTTGACATGGCAACAGAAGTTGGTTTCAGCATGCATCTGCTTGATATTGGTGGTGGCTTTCCTGG
ATCTGAAGATACAAAGCTTAAATTTGAAGAGATCACCAGTGTAATCAAACCAGCTCTGGACAAGTACTTCCCATCAGACT
CTGGAGTGAGAATCATAGCCGAGCCAGGCAGCTTTCACGCTTGCAGTCAACATCATTGCCAAAAAAAAAAAAAAAAAAAA
AAAAACGTGTGAAAGGAGCAGCCGAGTTCTGACGATGATGATGAGTCAAATGAGCAAACCTTCATGTATTATGTGAATGA
TGGAGTGTACGGATCATTTAACTGCATTCTTTATGACCATGCCCATGTGAAGGCCCTGCTGCAGAAGAGACCCAAGCCAG
ATGAGAAGTATTACTCATCCAGCATCTGGGGACCAACATGTGATGGCCTTGATCGGATCCTGGAGCGCTGTAACCTGCCT
GAAATGCATGTGGGTGATTGGATGCTGTTTGAGAACATGGGTGCATACACCGTTGCTGCTGCTTCTACTTTCAATGGGTT
CCAGAGGCCAAACATCTACTATGTAATGTCACGGCCAATGTGGCAACTCATGAAACAGATCCAGAGCCATGGCTTCCCGC
CGGAGGTGGAGGAGCAGGATGATGGCACGCTGCCCATGTCTTGTGCCCAGGAGAGCGGGATGGACCGTCACCCTGCAGCC
TGTGCTTCTGCTAGGATCAATGTG
ORF - retro_mmus_2588 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 92.64 % |
| Parental protein coverage: | 100. % |
| Number of stop codons detected: | 1 |
| Number of frameshifts detected | 1 |
Retrocopy - Parental Gene Alignment:
| Parental | MSSFTKDEFDCHILDEGFTAKDILDQK-INEVSSSDDKDAFYVADLGDILKKHLRWLKALPRVTPFYAVK |
| MS.FTKDEFDCHILDEGFTAKDIL.QK.INEVSSSDDKDAFYVADLGDILKKHLRWLKALPRVTP.YAVK | |
| Retrocopy | MSNFTKDEFDCHILDEGFTAKDILEQK>INEVSSSDDKDAFYVADLGDILKKHLRWLKALPRVTPVYAVK |
| Parental | CNDSRAIVSTLAAIGTGFDCASKTEIQLVQGLGVPAERVIYANPCKQVSQIKYAASNGVQMMTFDSEIEL |
| CNDSRAIVSTLAA.G.GFDCASKTEIQLVQG.GVP.E..IYANPCKQVSQIKYAAS.GVQMMTFDSEIEL | |
| Retrocopy | CNDSRAIVSTLAAFGIGFDCASKTEIQLVQGPGVPPETIIYANPCKQVSQIKYAASSGVQMMTFDSEIEL |
| Parental | MKVARAHPKAKLVLRIATDDSKAVCRLSVKFGATLKTSRLLLERAKELNIDVIGVSFHVGSGCTDPETFV |
| MKVARAHPKAKLVLRIATDDSKAVCRLSVKFGATLKTSRLLLERAKELNIDVIGVSFHVGSGCTDPETFV | |
| Retrocopy | MKVARAHPKAKLVLRIATDDSKAVCRLSVKFGATLKTSRLLLERAKELNIDVIGVSFHVGSGCTDPETFV |
| Parental | QAVSDARCVFDMATEVGFSMHLLDIGGGFPGSEDTKLKFEEITSVINPALDKYFPSDSGVRIIAEPGRYY |
| QAVSDARCVFDMATEVGFSMHLLDIGGGFPGSEDTKLKFEEITSVI.PALDKYFPSDSGVRIIAEPG... | |
| Retrocopy | QAVSDARCVFDMATEVGFSMHLLDIGGGFPGSEDTKLKFEEITSVIKPALDKYFPSDSGVRIIAEPGSFH |
| Parental | VASAFTLAVNIIAKKTVWKEQPGSDDEDESNEQTFMYYVNDGVYGSFNCILYDHAHVKALLQKRPKPDEK |
| ..S.............V.KEQP.SDD.DESNEQTFMYYVNDGVYGSFNCILYDHAHVKALLQKRPKPDEK | |
| Retrocopy | ACSQHHCXXXXXXXXXV*KEQPSSDDDDESNEQTFMYYVNDGVYGSFNCILYDHAHVKALLQKRPKPDEK |
| Parental | YYSSSIWGPTCDGLDRIVERCNLPEMHVGDWMLFENMGAYTVAAASTFNGFQRPNIYYVMSRPMWQLMKQ |
| YYSSSIWGPTCDGLDRI.ERCNLPEMHVGDWMLFENMGAYTVAAASTFNGFQRPNIYYVMSRPMWQLMKQ | |
| Retrocopy | YYSSSIWGPTCDGLDRILERCNLPEMHVGDWMLFENMGAYTVAAASTFNGFQRPNIYYVMSRPMWQLMKQ |
| Parental | IQSHGFPPEVEEQDDGTLPMSCAQESGMDRHPAACASARINV |
| IQSHGFPPEVEEQDDGTLPMSCAQESGMDRHPAACASARINV | |
| Retrocopy | IQSHGFPPEVEEQDDGTLPMSCAQESGMDRHPAACASARINV |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 0 .00 RPM | 36 .54 RPM |
| SRP007412_cerebellum | 0 .04 RPM | 43 .94 RPM |
| SRP007412_heart | 0 .12 RPM | 40 .72 RPM |
| SRP007412_kidney | 0 .42 RPM | 253 .38 RPM |
| SRP007412_liver | 0 .05 RPM | 51 .94 RPM |
| SRP007412_testis | 0 .50 RPM | 221 .74 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_2588 was not detected
No EST(s) were mapped for retro_mmus_2588 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_116676 | 560 libraries | 289 libraries | 195 libraries | 21 libraries | 7 libraries |
The graphical summary, for retro_mmus_2588 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
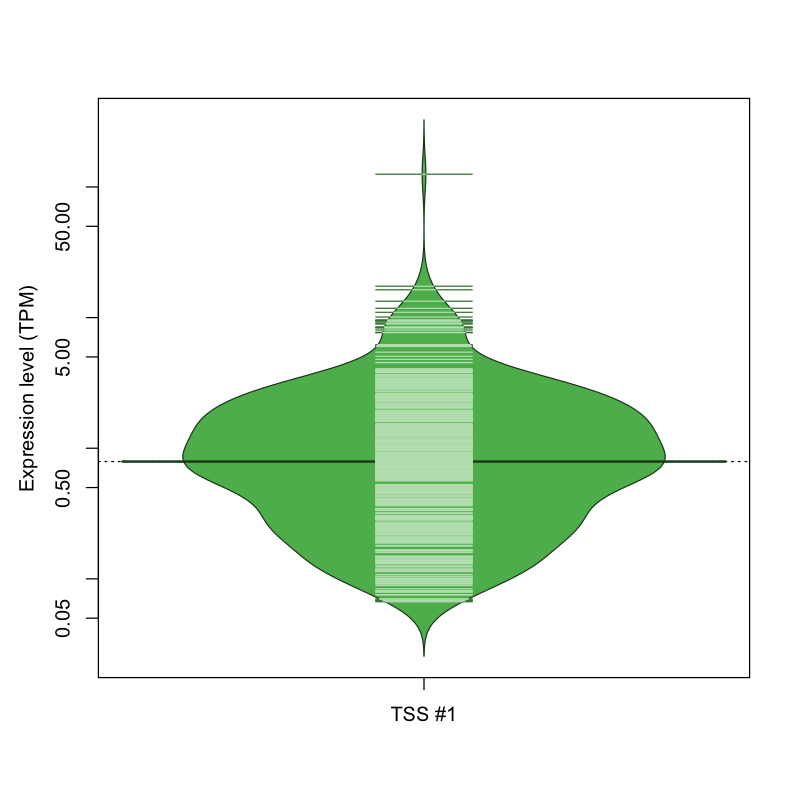
retro_mmus_2588 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_2588 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 8 parental genes, and 51 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Ailuropoda melanoleuca | ENSAMEG00000011415 | 1 retrocopy | |
| Bos taurus | ENSBTAG00000004256 | 1 retrocopy | |
| Dasypus novemcinctus | ENSDNOG00000018474 | 31 retrocopies |
retro_dnov_1032, retro_dnov_1128, retro_dnov_1129, retro_dnov_1137, retro_dnov_1212, retro_dnov_1235, retro_dnov_1418, retro_dnov_1434, retro_dnov_1493, retro_dnov_1594, retro_dnov_1595, retro_dnov_1677, retro_dnov_1778, retro_dnov_1810, retro_dnov_1813, retro_dnov_2016, retro_dnov_2027, retro_dnov_2108, retro_dnov_2147, retro_dnov_2180, retro_dnov_2212, retro_dnov_2461, retro_dnov_2551, retro_dnov_2564, retro_dnov_392, retro_dnov_740, retro_dnov_766, retro_dnov_781, retro_dnov_923, retro_dnov_943, retro_dnov_958,
|
| Myotis lucifugus | ENSMLUG00000010423 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000020441 | 2 retrocopies | |
| Monodelphis domestica | ENSMODG00000014779 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000011179 | 12 retrocopies | |
| Ictidomys tridecemlineatus | ENSSTOG00000000386 | 2 retrocopies |