RetrogeneDB ID: | retro_hsap_3313 | ||
Retrocopy location | Organism: | Human (Homo sapiens) | |
| Coordinates: | 5:43231916..43233275(-) | ||
| Located in intron of: | ENSG00000177453 | ||
Retrocopy information | Ensembl ID: | ENSG00000247763 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental gene information | Parental gene summary: | ||
| Parental gene symbol: | TUBA1A | ||
| Ensembl ID: | ENSG00000167552 | ||
| Aliases: | TUBA1A, B-ALPHA-1, LIS3, TUBA3 | ||
| Description: | tubulin, alpha 1a [Source:HGNC Symbol;Acc:20766] |

Retrocopy-Parental alignment summary:
>retro_hsap_3313
ATGCGTGACCGCATCTCCATCTCATTGGCCAGGCTGGTGTCCAGGTTGGCAATGCCTGCTGGGAGCCCTACTGCCTGGAA
CATGGCATCCAGCCCAATGGCCAGATGCCAAGTGACAAGACCACTCGGGGAGGAGATGACTCCTTCAACAGCTTCTTCAG
TGAGACGGGCTCCGGCAAGCATGTGCCCAGGGCAGTGTTTCTAGACCTGAAACCCATGGTAATTGATGATGAAGTTTGCA
CTGGCACCTACAACCAGCTCTTCCACCCTGAGCAGCTCATCACAGGCAAGGAAGATGCTTCCAGTAACTATGCCTGAAGG
CACTACACCATTGGCCAGGAGATCACTGACCTCGTTTTGGACCAAATTTGTAAACTGGCTGAACAGTGCATGGATCGCCA
AGGCTTCTTGTTTTTCCACAGCTTTGGTGAAGGAGTTGGTTCTGGGTTCACCTCCCCGCTCATGGAACAACTCCCGGTTG
ATTACGGCAAGAAGTCCAAGCTGCAGTTCTCCATTTACTCACCACCCCAGGTTTCCACAGCTGTAGTTGAGCCCTACAAC
TCCATCCTCACCTCCCACACCACCCTGGAGCACTCTGATTGTGTCTTCATGGTAGACAATGAGGCCACTTATGATATCTG
GCATAGAACCTTGATACTGAACGCCCAACCTATACTAAACTGAATAGGTTGATAGGTCAAATTGTGTCCTCCATCACTGC
TTCCCTGAGATTTGATGGAGTCCTGAATGTTGATCTGACAGAATTCCAGACCAAGCTGGTGGCCTATCCCCCCATCCACT
TCCGTCTGGACACATATGCCCTGTCATCTCTGCTGAGAAAGCCTACCATGAAACAGCTTTCTGTAGCAGAGACTGCCAGT
GCTTGCTTTCAGCCTGCCAACCAGATGATGAAATGTGAGCCTTGCCATGGTAAATACATGGCTTGCTGCCTGTTTTACCG
TGGTGACATGGTTCTCAAAGACGTCAATGCTGCCATTGCCACCATCAATCAAGACCAAGTGTATCATCTAATTTGTGGTT
TGGTGCCCCCTGGCTTCAAGGTTGGCATTAACTGCCAGCCCTCCCACTGTGGTGACTGGTGGAGACCTGGTCAAGACACA
GAGAGCTGTGTGCATGCTGAGCAATACCACAGCCATTGCTGAGGCCTGGGCTCACCTGGACCACAAGTTTGACTTGAATG
TATGTCAAGAGTGCTTTTGTTCACTGGTACGTGGGTGAGAGAAGGGAGGAAGGAGAGTTTTCTGAGGCCAGTGAGGACAT
GACTGCCCTTGAGAAGGATTATAAGGAGGTTGGTGTGGATTCTGTTGAAGGAGAAGGTGAGGAAGAAAGAGAGGAATAC
ORF - retro_hsap_3313 Open Reading Frame is not conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 79.52 % |
| Parental protein coverage: | 100.0 % |
| Number of stop codons detected: | 1 |
| Number of frameshifts detected: | 6 |
Retrocopy - Parental Gene Alignment:
| Parental | MRECISIH-VGQAGVQIGNACWELYCLEHGIQPDGQMPSDKTIGGGDDSFNTFFSETGAGKHVPRAVFVD |
| MR..ISI...GQAGVQ.GNACWE.YCLEHGIQP.GQMPSDKT..GGDDSFN.FFSETG.GKHVPRAVF.D | |
| Retrocopy | MRDRISIS<IGQAGVQVGNACWEPYCLEHGIQPNGQMPSDKTTRGGDDSFNSFFSETGSGKHVPRAVFLD |
| Parental | LEPTVID-EVRTGTYRQLFHPEQLITGKEDAANNYARGHYTIGKEIIDLVLDRIRKLADQCTGLQGFLVF |
| L.P.VID.EV.TGTY.QLFHPEQLITGKEDA..NYA..HYTIG.EI.DLVLD.I.KLA.QC...QGFL.F | |
| Retrocopy | LKPMVIDDEVCTGTYNQLFHPEQLITGKEDASSNYA*RHYTIGQEITDLVLDQICKLAEQCMDRQGFLFF |
| Parental | HSFGGGTGSGFTSLLMERLSVDYGKKSKLEFSIYPAPQVSTAVVEPYNSILTTHTTLEHSDCAFMVDNEA |
| HSFG.G.GSGFTS.LME.L.VDYGKKSKL.FSIY..PQVSTAVVEPYNSILT.HTTLEHSDC.FMVDNEA | |
| Retrocopy | HSFGEGVGSGFTSPLMEQLPVDYGKKSKLQFSIYSPPQVSTAVVEPYNSILTSHTTLEHSDCVFMVDNEA |
| Parental | IYDICRR-NLDIERPTYTNLNRLIGQIVSSITASLRFDGALNVDLTEFQTNLVPYPRIHFPLATYA-PVI |
| .YDI..R.NLD.ERPTYT.LNRLIGQIVSSITASLRFDG.LNVDLTEFQT.LV.YP.IHF.L.TYA.PVI | |
| Retrocopy | TYDIWHR<NLDTERPTYTKLNRLIGQIVSSITASLRFDGVLNVDLTEFQTKLVAYPPIHFRLDTYA<PVI |
| Parental | SAEKAYHE-QLSVAEITNACFEPANQMVKCDPRHGKYMACCLLYRGDVVPKDVNAAIATIKTKRTIQFV- |
| SAEKAYHE.QLSVAE...ACF.PANQM.KC.P.HGKYMACCL.YRGD.V.KDVNAAIATI.......... | |
| Retrocopy | SAEKAYHE>QLSVAETASACFQPANQMMKCEPCHGKYMACCLFYRGDMVLKDVNAAIATINQDQVYHLIC |
| Parental | DWCPTGFKVGINYQ-PPTVVPGGDLAKVQRAVCMLSNTTAIAEAWARLDHKFDL-MYAKRAFVHWYVGEG |
| ...P.GFKVGIN.Q.PPTVV.GGDL.K.QRAVCMLSNTTAIAEAWA.LDHKFDL.MY.K.AFVHWYVGE. | |
| Retrocopy | GLVPPGFKVGINCQ>PPTVVTGGDLVKTQRAVCMLSNTTAIAEAWAHLDHKFDL>MYVKSAFVHWYVGER |
| Parental | MEEGEFSEAREDMAALEKDYEEVGVDSVEGEGEEEGEEY |
| .EEGEFSEA.EDM.ALEKDY.EVGVDSVEGEGEEE.EEY | |
| Retrocopy | REEGEFSEASEDMTALEKDYKEVGVDSVEGEGEEEREEY |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| bodymap2_adipose | 0 .00 RPM | 40 .79 RPM |
| bodymap2_adrenal | 0 .00 RPM | 138 .19 RPM |
| bodymap2_brain | 0 .07 RPM | 685 .29 RPM |
| bodymap2_breast | 0 .00 RPM | 120 .64 RPM |
| bodymap2_colon | 0 .00 RPM | 124 .87 RPM |
| bodymap2_heart | 0 .04 RPM | 57 .28 RPM |
| bodymap2_kidney | 0 .00 RPM | 99 .27 RPM |
| bodymap2_liver | 0 .00 RPM | 5 .58 RPM |
| bodymap2_lung | 0 .00 RPM | 83 .42 RPM |
| bodymap2_lymph_node | 0 .00 RPM | 60 .81 RPM |
| bodymap2_ovary | 0 .06 RPM | 189 .07 RPM |
| bodymap2_prostate | 0 .00 RPM | 127 .93 RPM |
| bodymap2_skeletal_muscle | 0 .00 RPM | 10 .97 RPM |
| bodymap2_testis | 0 .00 RPM | 88 .18 RPM |
| bodymap2_thyroid | 0 .00 RPM | 23 .80 RPM |
| bodymap2_white_blood_cells | 0 .00 RPM | 180 .52 RPM |
RNA Polymerase II activity near the 5' end of retro_hsap_3313 was not detected
No EST(s) were mapped for retro_hsap_3313 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_154282 | 1047 libraries | 613 libraries | 167 libraries | 2 libraries | 0 libraries |
The graphical summary, for retro_hsap_3313 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1829 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
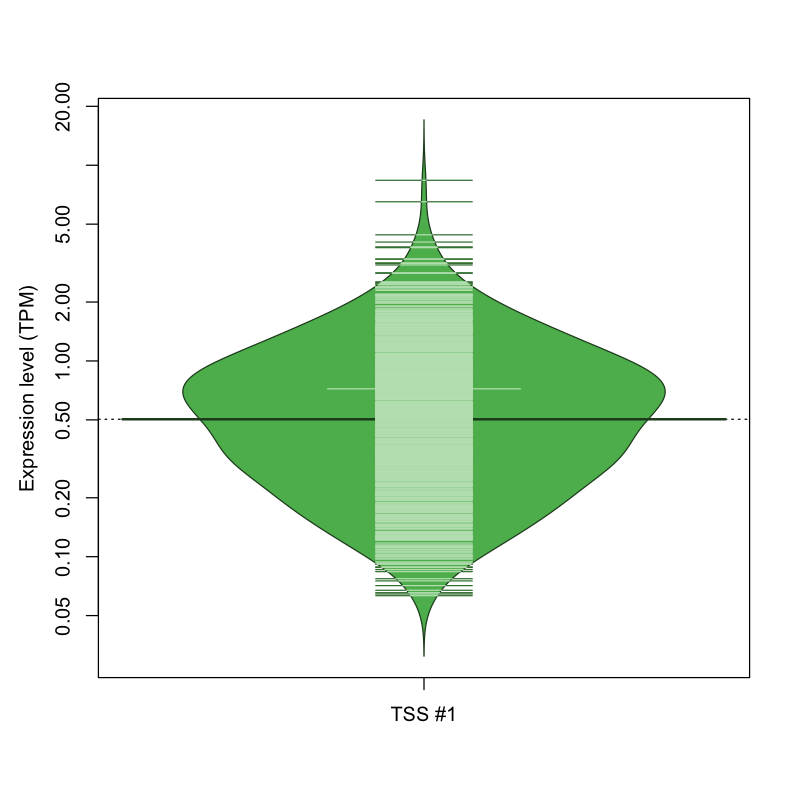
retro_hsap_3313 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_hsap_3313 has 1 orthologous retrocopies within eutheria group .| Species | RetrogeneDB ID |
|---|---|
| Pan troglodytes | retro_ptro_2158 |
Parental genes homology:
Parental genes homology involve 11 parental genes, and 21 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Bos taurus | ENSBTAG00000001489 | 3 retrocopies | |
| Callithrix jacchus | ENSCJAG00000020894 | 2 retrocopies | |
| Dasypus novemcinctus | ENSDNOG00000004121 | 1 retrocopy | |
| Equus caballus | ENSECAG00000013798 | 1 retrocopy | |
| Homo sapiens | ENSG00000123416 | 1 retrocopy | |
| Homo sapiens | ENSG00000167552 | 1 retrocopy |
retro_hsap_3313 ,
|
| Nomascus leucogenys | ENSNLEG00000017836 | 2 retrocopies | |
| Oryctolagus cuniculus | ENSOCUG00000027812 | 1 retrocopy | |
| Pteropus vampyrus | ENSPVAG00000004645 | 2 retrocopies | |
| Rattus norvegicus | ENSRNOG00000004559 | 3 retrocopies | |
| Vicugna pacos | ENSVPAG00000005749 | 4 retrocopies |
Expression level across human populations :
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2089
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2089
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2112
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2112
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2135
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2135
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2158
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2158
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2181
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2181
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2204
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2204
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2227
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2227
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2250
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2250
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2273
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2273
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2296
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2296
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2325
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2325
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2348
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2348
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2371
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2371
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2394
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2394
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2417
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2417
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2440
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2440
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2463
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2463
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2486
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2486
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2509
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2509
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2532
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2532
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2553
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2553
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2576
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2576
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2599
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2599
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2622
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2622
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2645
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2645
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2668
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2668
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2691
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2691
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2714
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2714
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2737
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2737
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2760
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2760
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2787
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2787
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2810
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2810
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2837
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2837
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2860
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2860
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2887
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2887
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2910
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2910
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2937
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2937
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2960
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2960
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2987
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 2987
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3010
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3010
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3122
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3122
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3145
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3145
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3168
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3168
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3191
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3191
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3214
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3214
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3241
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3241
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3264
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3264
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3287
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3287
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3310
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3310
Warning: Undefined variable $populationsDictCols in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3333
Warning: Trying to access array offset on value of type null in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3333
Warning: Undefined variable $populLEGEND in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3353
| Library | Retrogene expression |
|---|---|
| CEU_NA11831 | 0 .00 RPM |
| CEU_NA11843 | 0 .00 RPM |
| CEU_NA11930 | 0 .00 RPM |
| CEU_NA12004 | 0 .00 RPM |
| CEU_NA12400 | 0 .00 RPM |
| CEU_NA12751 | 0 .00 RPM |
| CEU_NA12760 | 0 .00 RPM |
| CEU_NA12827 | 0 .00 RPM |
| CEU_NA12872 | 0 .00 RPM |
| CEU_NA12873 | 0 .00 RPM |
| FIN_HG00183 | 0 .00 RPM |
| FIN_HG00277 | 0 .00 RPM |
| FIN_HG00315 | 0 .00 RPM |
| FIN_HG00321 | 0 .00 RPM |
| FIN_HG00328 | 0 .00 RPM |
| FIN_HG00338 | 0 .00 RPM |
| FIN_HG00349 | 0 .00 RPM |
| FIN_HG00375 | 0 .00 RPM |
| FIN_HG00377 | 0 .00 RPM |
| FIN_HG00378 | 0 .00 RPM |
| GBR_HG00099 | 0 .00 RPM |
| GBR_HG00111 | 0 .00 RPM |
| GBR_HG00114 | 0 .00 RPM |
| GBR_HG00119 | 0 .00 RPM |
| GBR_HG00131 | 0 .00 RPM |
| GBR_HG00133 | 0 .00 RPM |
| GBR_HG00134 | 0 .00 RPM |
| GBR_HG00137 | 0 .00 RPM |
| GBR_HG00142 | 0 .00 RPM |
| GBR_HG00143 | 0 .00 RPM |
| TSI_NA20512 | 0 .00 RPM |
| TSI_NA20513 | 0 .00 RPM |
| TSI_NA20518 | 0 .00 RPM |
| TSI_NA20532 | 0 .00 RPM |
| TSI_NA20538 | 0 .00 RPM |
| TSI_NA20756 | 0 .00 RPM |
| TSI_NA20765 | 0 .00 RPM |
| TSI_NA20771 | 0 .00 RPM |
| TSI_NA20786 | 0 .00 RPM |
| TSI_NA20798 | 0 .00 RPM |
| YRI_NA18870 | 0 .00 RPM |
| YRI_NA18907 | 0 .00 RPM |
| YRI_NA18916 | 0 .00 RPM |
| YRI_NA19093 | 0 .00 RPM |
| YRI_NA19099 | 0 .00 RPM |
| YRI_NA19114 | 0 .00 RPM |
| YRI_NA19118 | 0 .00 RPM |
| YRI_NA19213 | 0 .00 RPM |
| YRI_NA19214 | 0 .00 RPM |
| YRI_NA19223 | 0 .00 RPM |
Fatal error: Uncaught Error: Call to undefined function mysql_query() in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php:3390 Stack trace: #0 /home/rosikiewicz/RetrogeneDB2/BETA/retrogene.php(1158): include() #1 {main} thrown in /home/rosikiewicz/RetrogeneDB2/BETA/retrogene_maps.php on line 3390