RetrogeneDB ID: | retro_mmus_1817 | ||
Retrocopylocation | Organism: | Mouse (Mus musculus) | |
| Coordinates: | 19:21937379..21938381(+) | ||
| Located in intron of: | None | ||
Retrocopyinformation | Ensembl ID: | ENSMUSG00000045104 | |
| Aliases: | None | ||
| Status: | KNOWN_PSEUDOGENE | ||
Parental geneinformation | Parental gene summary: | ||
| Parental gene symbol: | Ldhb | ||
| Ensembl ID: | ENSMUSG00000030246 | ||
| Aliases: | Ldhb, AI790582, H-Ldh, LDH-B, LDH-H, Ldh-2, Ldh2 | ||
| Description: | lactate dehydrogenase B [Source:MGI Symbol;Acc:MGI:96763] |

Retrocopy-Parental alignment summary:
>retro_mmus_1817
ATGGCAACCCTTAAGGAGAAGCTCATTGCATCAGTTGCAGAAGATGAGGCTGCCGTCCCGAACAACAAGATCACTGTAGT
GGGCGTTGGACAAGTGGGTATGGCGTGTGCCATCAGCATTCTGGGAAAGTCTCTGGCTGATGAACTTGCTCTGGTGGATG
TGTTGGAAGACAAGCTCAAAGAAGAGATGATGGACCTGCAGCACGGGAGCTTGTTCCTCCAGACTCCGAAAATTGTGGCC
GATAAAGATTACTCTGTGACAGCCAACTCTAAGATTGTGGTGGTGTCCGCAGGAGTGCGCCAGCAGGAGGGGGAGAGTCG
GCTCAACCTGGTGCAGAGAAATGTCAATGTGTTCAAGTTCATCATTCCTCAGATCGTCAAATACAGCCCTGACTGCACCA
TCATCGTGGTTTCCAACCCAGTGGATATTCTGACTTATGTCACCTGGAAACTGAGAGGGCTACCTAAGCACCGTGTGATT
GGAAGCGGATGCAATCTGGATTCTGCTCGATTCCGCTACCTCATGGCAGAGAAGCTTGGCATTCATCCCAGCAGCTGCCA
TGGATGGATCCTGGGCGAGCATGGAGACTCCAGTGTGGCTGTGTGGAGCGGGGTGAATGTGGCAGGAGTCTCCCTCCAGG
AACTGAATCCAGAAATGGGGACAGACAATGACAGTGAGAACTGGAAGGAGGTGCATAAGATGGTGGTGGTCAGTGCCTAT
GAAGTCATCAAGCTCAAAGGCTACACCAACTGGGCCATCGGCCTGAGCGTGGCTGACCTCATCGAGTCCATGCTGAAAAA
CCTCTCCCGGATTCACCCCGTGTCTACCATGGTGAAGGGAATGTACGGCATTGAGAACGAAGTCTTCCTCAGTCTCCCAT
GTATCCTCAACGCTCGGGGGCTGACCAGCGTCATCAATCAGAAGCTGAAGGACGATGAGGTTGCTCAGCTCAGGAAGAGT
GCGGACACCCTGTGGGACATCCAGAAAGACCTCAAAGACCTG
ATGGCAACCCTTAAGGAGAAGCTCATTGCATCAGTTGCAGAAGATGAGGCTGCCGTCCCGAACAACAAGATCACTGTAGT
GGGCGTTGGACAAGTGGGTATGGCGTGTGCCATCAGCATTCTGGGAAAGTCTCTGGCTGATGAACTTGCTCTGGTGGATG
TGTTGGAAGACAAGCTCAAAGAAGAGATGATGGACCTGCAGCACGGGAGCTTGTTCCTCCAGACTCCGAAAATTGTGGCC
GATAAAGATTACTCTGTGACAGCCAACTCTAAGATTGTGGTGGTGTCCGCAGGAGTGCGCCAGCAGGAGGGGGAGAGTCG
GCTCAACCTGGTGCAGAGAAATGTCAATGTGTTCAAGTTCATCATTCCTCAGATCGTCAAATACAGCCCTGACTGCACCA
TCATCGTGGTTTCCAACCCAGTGGATATTCTGACTTATGTCACCTGGAAACTGAGAGGGCTACCTAAGCACCGTGTGATT
GGAAGCGGATGCAATCTGGATTCTGCTCGATTCCGCTACCTCATGGCAGAGAAGCTTGGCATTCATCCCAGCAGCTGCCA
TGGATGGATCCTGGGCGAGCATGGAGACTCCAGTGTGGCTGTGTGGAGCGGGGTGAATGTGGCAGGAGTCTCCCTCCAGG
AACTGAATCCAGAAATGGGGACAGACAATGACAGTGAGAACTGGAAGGAGGTGCATAAGATGGTGGTGGTCAGTGCCTAT
GAAGTCATCAAGCTCAAAGGCTACACCAACTGGGCCATCGGCCTGAGCGTGGCTGACCTCATCGAGTCCATGCTGAAAAA
CCTCTCCCGGATTCACCCCGTGTCTACCATGGTGAAGGGAATGTACGGCATTGAGAACGAAGTCTTCCTCAGTCTCCCAT
GTATCCTCAACGCTCGGGGGCTGACCAGCGTCATCAATCAGAAGCTGAAGGACGATGAGGTTGCTCAGCTCAGGAAGAGT
GCGGACACCCTGTGGGACATCCAGAAAGACCTCAAAGACCTG
ORF - retro_mmus_1817 Open Reading Frame is conserved.
Retrocopy - Parental Gene Alignment summary:
| Percent Identity: | 98.5 % |
| Parental protein coverage: | 100. % |
| Number of stop codons detected: | 0 |
| Number of frameshifts detected | 0 |
Retrocopy - Parental Gene Alignment:
| Parental | MATLKEKLIASVADDEAAVPNNKITVVGVGQVGMACAISILGKSLADELALVDVLEDKLKGEMMDLQHGS |
| MATLKEKLIASVA.DEAAVPNNKITVVGVGQVGMACAISILGKSLADELALVDVLEDKLK.EMMDLQHGS | |
| Retrocopy | MATLKEKLIASVAEDEAAVPNNKITVVGVGQVGMACAISILGKSLADELALVDVLEDKLKEEMMDLQHGS |
| Parental | LFLQTPKIVADKDYSVTANSKIVVVTAGVRQQEGESRLNLVQRNVNVFKFIIPQIVKYSPDCTIIVVSNP |
| LFLQTPKIVADKDYSVTANSKIVVV.AGVRQQEGESRLNLVQRNVNVFKFIIPQIVKYSPDCTIIVVSNP | |
| Retrocopy | LFLQTPKIVADKDYSVTANSKIVVVSAGVRQQEGESRLNLVQRNVNVFKFIIPQIVKYSPDCTIIVVSNP |
| Parental | VDILTYVTWKLSGLPKHRVIGSGCNLDSARFRYLMAEKLGIHPSSCHGWILGEHGDSSVAVWSGVNVAGV |
| VDILTYVTWKL.GLPKHRVIGSGCNLDSARFRYLMAEKLGIHPSSCHGWILGEHGDSSVAVWSGVNVAGV | |
| Retrocopy | VDILTYVTWKLRGLPKHRVIGSGCNLDSARFRYLMAEKLGIHPSSCHGWILGEHGDSSVAVWSGVNVAGV |
| Parental | SLQELNPEMGTDNDSENWKEVHKMVVDSAYEVIKLKGYTNWAIGLSVADLIESMLKNLSRIHPVSTMVKG |
| SLQELNPEMGTDNDSENWKEVHKMVV.SAYEVIKLKGYTNWAIGLSVADLIESMLKNLSRIHPVSTMVKG | |
| Retrocopy | SLQELNPEMGTDNDSENWKEVHKMVVVSAYEVIKLKGYTNWAIGLSVADLIESMLKNLSRIHPVSTMVKG |
| Parental | MYGIENEVFLSLPCILNARGLTSVINQKLKDDEVAQLRKSADTLWDIQKDLKDL |
| MYGIENEVFLSLPCILNARGLTSVINQKLKDDEVAQLRKSADTLWDIQKDLKDL | |
| Retrocopy | MYGIENEVFLSLPCILNARGLTSVINQKLKDDEVAQLRKSADTLWDIQKDLKDL |
Legend:
| * | Stop codon |
| > | Forward frameshift by one nucleotide |
| < | Reverse frameshift by one nucleotide |
(Hint: click retrocopy or parental gene accession number on the plot's legend, to show / hide expression level values)
Expression validation based on RNA-Seq data:
| Library | Retrocopy expression | Parental gene expression |
|---|---|---|
| SRP007412_brain | 4 .98 RPM | 323 .16 RPM |
| SRP007412_cerebellum | 16 .93 RPM | 420 .62 RPM |
| SRP007412_heart | 2 .93 RPM | 1199 .98 RPM |
| SRP007412_kidney | 39 .56 RPM | 997 .84 RPM |
| SRP007412_liver | 0 .05 RPM | 2 .65 RPM |
| SRP007412_testis | 3 .11 RPM | 63 .67 RPM |
RNA Polymerase II actvity near the 5' end of retro_mmus_1817 was not detected
No EST(s) were mapped for retro_mmus_1817 retrocopy.
| TSS No. | TSS Name | TSS expression level (Expr) in TPM range: | ||||
|---|---|---|---|---|---|---|
| no expression | 0 < Expr ≤ 1 | 1 < Expr ≤ 5 | 5 < Expr ≤ 10 | Expr > 10 | ||
| TSS #1 | TSS_63039 | 494 libraries | 283 libraries | 184 libraries | 44 libraries | 67 libraries |
The graphical summary, for retro_mmus_1817 TSS expression levels > 0 TPM .
TSS expression levels were studied across 1072 TSS-CAGE libraries, based on FANTOM5 data.
The expression values were visualized using beanplot. If you have any doubts, how to read it, read more in Kampstra P (2008)
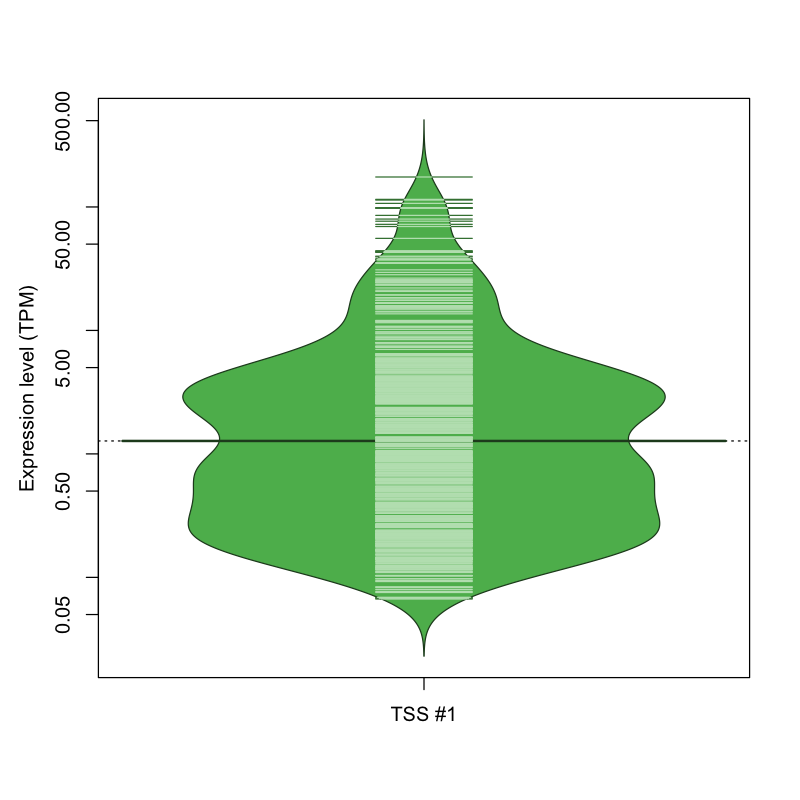
retro_mmus_1817 was not experimentally validated.
Retrocopy orthology:
Retrocopy retro_mmus_1817 has 0 orthologous retrocopies within eutheria group .Parental genes homology:
Parental genes homology involve 19 parental genes, and 66 retrocopies.| Species | Parental gene accession | Retrocopies number | |
|---|---|---|---|
| Ailuropoda melanoleuca | ENSAMEG00000016175 | 4 retrocopies | |
| Canis familiaris | ENSCAFG00000012195 | 9 retrocopies | |
| Cavia porcellus | ENSCPOG00000007138 | 1 retrocopy | |
| Dasypus novemcinctus | ENSDNOG00000013620 | 4 retrocopies | |
| Felis catus | ENSFCAG00000012639 | 1 retrocopy | |
| Gorilla gorilla | ENSGGOG00000015758 | 1 retrocopy | |
| Loxodonta africana | ENSLAFG00000000972 | 2 retrocopies | |
| Macropus eugenii | ENSMEUG00000004825 | 1 retrocopy | |
| Microcebus murinus | ENSMICG00000017183 | 2 retrocopies | |
| Myotis lucifugus | ENSMLUG00000012155 | 1 retrocopy | |
| Macaca mulatta | ENSMMUG00000002679 | 4 retrocopies | |
| Monodelphis domestica | ENSMODG00000017786 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000030246 | 4 retrocopies | |
| Mus musculus | ENSMUSG00000043262 | 1 retrocopy | |
| Mus musculus | ENSMUSG00000063229 | 14 retrocopies | |
| Otolemur garnettii | ENSOGAG00000004763 | 9 retrocopies | |
| Pteropus vampyrus | ENSPVAG00000002462 | 3 retrocopies | |
| Rattus norvegicus | ENSRNOG00000013000 | 3 retrocopies | |
| Sus scrofa | ENSSSCG00000000576 | 1 retrocopy |